This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1dfu
From Proteopedia
|
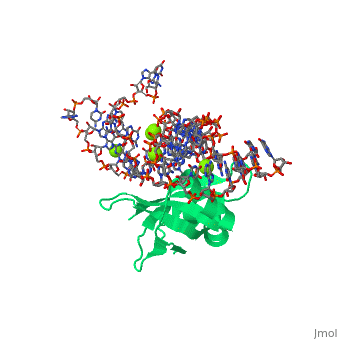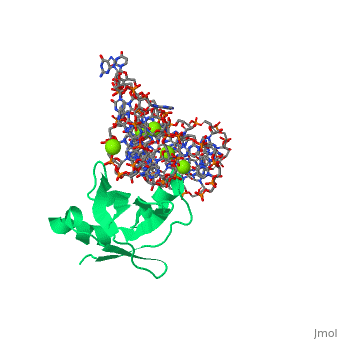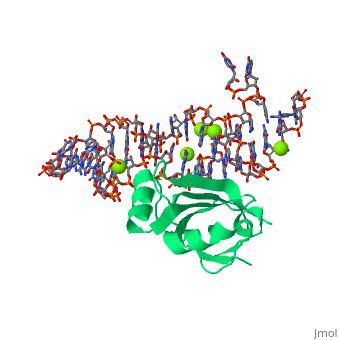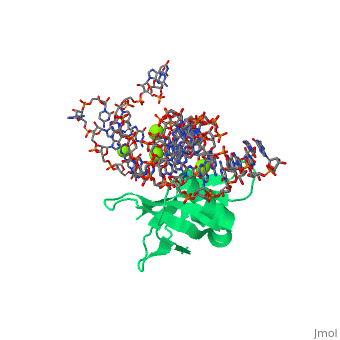
CRYSTAL STRUCTURE OF E.COLI RIBOSOMAL PROTEIN L25 COMPLEXED WITH A 5S RRNA FRAGMENT AT 1.8 A RESOLUTION
Overview
The crystal structure of Escherichia coli ribosomal protein L25 bound to, an 18-base pair portion of 5S ribosomal RNA, which contains "loop E," has, been determined at 1.8-A resolution. The protein primarily recognizes a, unique RNA shape, although five side chains make direct or water-mediated, interactions with bases. Three beta-strands lie in the widened minor, groove of loop E formed by noncanonical base pairs and cross-strand purine, stacks, and an alpha-helix interacts in an adjacent widened major groove., The structure of loop E is largely the same as that of uncomplexed RNA, (rms deviation of 0.4 A for 11 base pairs), and 3 Mg(2+) ions that, stabilize the noncanonical base pairs lie in the same or similar locations, in both structures. Perhaps surprisingly, those residues interacting with, the RNA backbone are the most conserved among known L25 sequences, whereas, those interacting with the bases are not.
About this Structure
1DFU is a Single protein structure of sequence from Escherichia coli with MG as ligand. Full crystallographic information is available from OCA.
Reference
Structure of Escherichia coli ribosomal protein L25 complexed with a 5S rRNA fragment at 1.8-A resolution., Lu M, Steitz TA, Proc Natl Acad Sci U S A. 2000 Feb 29;97(5):2023-8. PMID:10696113
Page seeded by OCA on Tue Nov 20 13:15:23 2007