This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
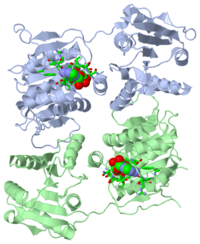
DEAD-box RNA-helicase DDX19 in complex with ADP
From Proteopedia
| |||||||||||
Reference
- Collins R, Karlberg T, Lehtio L, Schutz P, van den Berg S, Dahlgren LG, Hammarstrom M, Weigelt J, Schuler H. The DEXD/H-box RNA helicase DDX19 is regulated by an {alpha}-helical switch. J Biol Chem. 2009 Apr 17;284(16):10296-300. Epub 2009 Feb 25. PMID:19244245 doi:10.1074/jbc.C900018200
Created with the participation of Wayne Decatur.
Categories: Homo sapiens | Andersson,j. | Arrowsmith,c h. | Berglund,h. | Bountra, c. | Collins,r. | Dahlgren,l g. | Edwards,a m. | Flodin,s. | Flores,a. | Graslund,s. | Hammarstrom,m. | Johansson,a. | Johansson,i. | Karlberg,t. | Kotenyova,t. | Lehtio,l. | Moche,m. | Nilsson,m e. | Nordlund,p. | Nyman,t. | Olesen,k. | Persson,c. | SGC, Structural Genomics Consortium. | Sagemark,j. | Schueler,h. | Thorsell,a g. | Tresaugues,l. | Weigelt, j. | Welin,m. | Wikstrom,m. | Wisniewska,m. | Berg, s Van den. | Adp | Alternative splicing | Atp-binding | Atpase | Cytoplasm | Dead | Helicase | Hydrolase | Membrane | Mrna | Mrna transport | Nuclear pore complex | Nucleotide-binding | Nucleus | Protein transport | Rna helicase | Rna-binding | Rrna | Sgc | Structural genomic | Structural genomics consortium | Translocation | Transport