This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Johnson's Monday Lab Sandbox for Insulin Receptor
From Proteopedia
Insulin Receptor
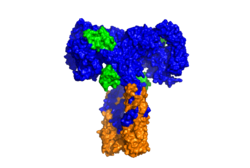
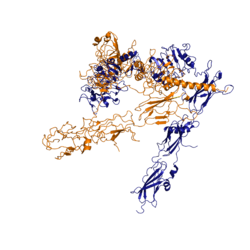
Function of the ReceptorThe insulin receptor binds the insulin hormone and initiates a cascade of events within the cell. The receptor resides within the plasma membrane of insulin targeted cells. These cells are found in various organs, such as the liver, and tissues, including skeletal muscle and adipose [1]. The insulin receptor is activated by multiple insulin molecules binding to various sites on the receptor [2]. Once activated, the receptor serves as the gateway for the regulation of various cellular processes including glucose transport, glycogen storage, autophagy, apoptosis, and gene expression. Additionally, problems with the insulin receptor are associated with the development of diseases such as Alzheimer's, type II diabetes, and cancer [3]. Recent structures of the insulin receptor have illustrated the large scale conformational changes, initiated by insulin binding. Evaluation of the structural composition and the biochemical properties of the insulin receptor reveals details about the role of the receptor in crucial cellular processes. InsulinThe is a hormone made of two separate amino acid chains that are bound by multiple disulfide bonds. Insulin is synthesized and secreted from the islets of Langerhans of the pancreas in response to high concentrations of glucose in the blood. Once it is secreted, insulin moves through the bloodstream and binds to unactivated insulin receptors residing in the plasma membrane. Binding of insulin to the insulin receptor is a complex process, which involves negative cooperativity among insulin molecules [2] [4] [5]. Current hypotheses propose that the receptor is fully activated only after multiple insulin molecules are bound [2]. StructureThe insulin receptor is a receptor tyrosine kinase. It is a heterotetramer that is constructed from two homodimers. Each homodimer maintains an extracellular domain, transmembrane helix, and an intracellular domain. The insulin receptor is divided into subunits. The alpha subunit is characterized by two leucine-rich regions and one cysteine-rich region. The beta subunit contains three fibronectin type III domains along with the transmembrane domain and intracellular tyrosine kinase domain that could not be shown in one continous PDB structure. The alpha and beta subunits of the extracellular domains fold over one another and form a when the insulin receptor is inactivated. Upon activation, the extracellular domain undergoes a conformational change and forms a . An additional component to the ectodomain is the , which is also referred to as the "alpha-CT" [2]. Each of the dimers has an "alpha"-CT. The alpha-CT is a single alpha-helix and it plays an important role in insulin binding and stabilization of the "T" shape activated conformation. The alpha-CT interacts with a leucine-rich region of the alpha subunit and a fibronectin type III region of the beta subunit to form the insulin binding sites known as [2]. The structure of the extracellular domain is stabilized through multiple disulfide bonds. The alpha subunits are linked through two disulfide bonds, with the main one being between of two adjacent alpha subuntis [4]. of both alpha subunits are also held together with a disulfide bond [6]. The alpha subunit is also attached to the beta subunit by a disulfide bond between the [6]. Insulin BindingThe insulin receptor unit has four separate sites for the insulin binding. There are two pairs of two identical binding sites referred to as and . The insulin molecules bind to these sites mostly through hydrophobic interactions, with some of the most crucial residues at sites 1 and 1' being between of the insulin receptor FnIII-1 domain [2]. Despite some of the residues included being charged they can still interact hydrophobically in this binding site. For example, due to arginine carrying its positive charge at the end of the side chain, to allow the hydrophobic part of the side chain to interact with the other hydrophobic residues. At sites 2 and 2', the major residues contributing to these hydrophobic interactions are the [2]. While the majority of the binding interactions appear similar, sites 1 and 1' have a higher binding affinity than sites 2 and 2' due to site 1 having a larger surface area (706 Å2) exposed for insulin to bind to compared to site 2 (394 Å2)[2]. The binding interactions of the insulin molecules in sites 1 and 1' are facilitated by hydrophobic residues of an of the insulin receptor. The insulin molecules in sites 2 and 2' primarily interact with the residues that comprise some of the of the insulin receptor. The secondary structures themselves are not what directly causes the differences in binding affinities, but the surface area that the insulin molecule can interact with. Recent studies have demonstrated that at least three insulin molecules have to bind to the insulin receptor to induce the active conformation, as binding of two insulin molecules is insufficient to induce a full conformational change [2]. However, this conclusion has not yet been widely confirmed [2]. It has been speculated that activation of the insulin receptor can change based on the concentration of insulin. In low concentrations of insulin, the insulin receptor may not require binding of three insulin molecules in order to exhibit activation. Rather, the level of activity will change in accordance to the availability of insulin [2]. When higher concentrations of insulin are present, the conformational difference between the two-insulin-bound state and the three-insulin-bound state is drastic as the insulin receptor transitions from the inactive to the active [2]. However, in conditions of low insulin availability, the two-insulin-bound state may be enough to induce partial activation of the receptor [2]. Conformational ChangesThe conformational change between the inverted, inactive and the active of the insulin receptor is induced by insulin binding. When an insulin molecule binds to site 1 of the alpha subunit, the respective protomer is recruited and a slight inward movement of the of the beta subunit is initiated. This is accomplished by the formation of several salt bridges, specifically between [2]. Binding of insulin to both protomers establishes a full activation of the insulin receptor. This activation is demonstrated through the inward movement of both protomers. This motion has been referred to as a "hinge" motion [2] as both protomers "swing" in towards one another. Figure 3 depicts the conformational change and "hinge motion" between the inactive and active forms of an insulin receptor protomer. Upon insulin binding, the beta subunits of the inactive form, shown in blue, are "swung" inward to the active form, shown in orange. As the fibronectin type III domains of the beta subunit swing inward, the alpha subunits also undergo a conformational change upon insulin binding. As insulin binds to site 1, the leucine-rich region of one protomer interacts with the alpha-CT and the FNIII-1 domains of the other protomer to form a binding site. These interactions are referred to as the [2]. In order for the tripartite interface to form, the alpha subunits of each protomer must undergo a "folding" motion. The proper conformational change of the ectodomain of the insulin receptor is crucial for transmitting the signal into the cell. The movements extracellularly cause the two receptor tyrosine kinase domains intracellularly to become close enough to each other to autophosphorylate [1]. This autophosphorylation leads enzymes to become activated in the cell that carries out processes related to insulin signaling such as metabolism and growth [1]. While there is an explanation for which conformational changes of the insulin receptor take place, there is no full explanation for the exact mechanism by which the conformational changes are executed in the receptor [2]. It is known where the various domains move, but not the specifics for how this is achieved on the atomic level due to the complexity of analyzing moving structures. Type II DiabetesType II diabetes (T2D) is a chronic condition that affects 10 percent of the world's population [1]. T2D is characterized by insulin resistance and leads to high concentrations of glucose in the bloodstream. A type II diabetic produces insulin, but when the insulin molecule binds to the insulin receptor, the signal is not properly transmitted intracellularly [1]. Insulin resistance in routine type II diabetics is not associated with mutations of the insulin receptor gene, but instead, the signal being disrupted later in the pathway [1]. Mutations of the receptor gene are associated with more severe cases of insulin resistance, as seen in leprechaunism. Additionally, mutations of the insulin receptor can be fatal, as it is crucial for many cellular processes including gene expression, glucose homeostasis, and apoptosis [1]. The basis for insulin resistance in typical type II diabetics is complex and cannot yet be explained by one particular factor [7] [1]. There are a multitude of hypotheses which discuss the reasons for the establishment of type II diabetes [1] [7]. While the specifics of the development of T2D are beyond the scope of this article, it is important to note that the molecular causes for insulin resistance and T2D have been primarily attributed to the inhibition of key proteins involved in the insulin signaling and glucose transport pathway [1]. Alterations to the phosphorylation cascade of insulin signaling can be the result of changes within the cellular environment including lipotoxicity, inflammation, hyperglycemia, and the presence of reactive oxygen species (ROS) [1]. On a macroscopic level, a variety of factors influence the cellular environment, and thus the risk for T2D. These factors include gestational environment, microbiome, genetics, diet, and energy expenditure [7]. Recent studies, which have evaluated the relationships between genetics and environmental factors in the progress of T2D, have shown that T2D is not uniform among the population and the biochemistry behind the development of risk factors varies for each patient [7].
References
Student Contributors
| ||||||||||||