This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox BCMB402 Ricin
From Proteopedia
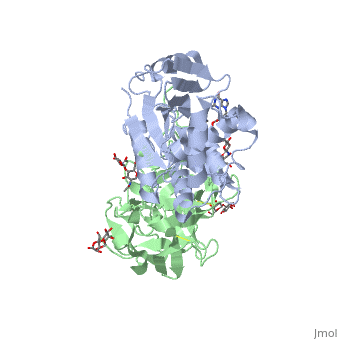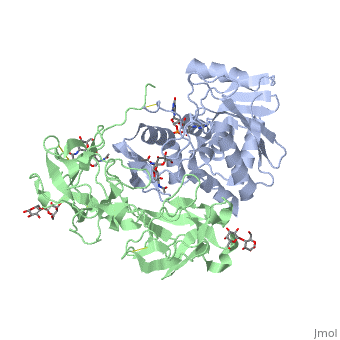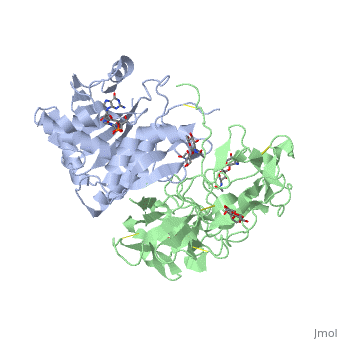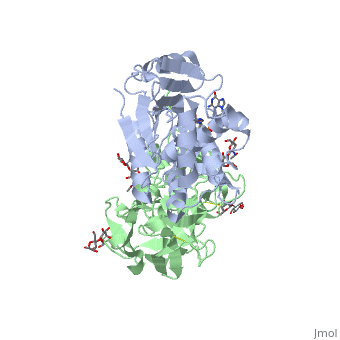
Ricin is a potent cytotoxin that is synthesized in the endosperm cells of maturing seeds of the castor oil plant (Ricinus communis)[1]. Ricin belongs to a small multi-gene family[2] that is composed of eight members. Ricin is classified as a type II heterodimeric Ribosome Inactivating Protein[1] or RIPs. For toxins in Proteopedia see Toxins.
| |||||||||||
Site of ricin modification of rRNA
| |||||||||||
Updated April 2013
Ricin A chain (RTA)
1j1m, 1ift, 2aai, 1rtc – RTA
3lc9, 3mk9, 2vc4, 1uq4, 1uq5, 1obs, 3bjg, 3srp – RTA (mutant)
Ricin A chain binary complexes
3px8 – RTA preproricin + 7-carboxy-pterin
1br5, 1br6 - RTA + pterin derivative
3px9 - RTA preproricin + furanylmethyl-carbamoyl-pterin
3lc9, 3mk9, 2vc4, 1uq4, 1uq5, 1obs – RTA (mutant)
3hio – RTA + tetranucleotide
3ej5, 1il5 – RTA pyrimidine derivative
2p8n, 1ifs – RTA + adenine
2pjo, 2r2x – RTA + urea derivative
2r3d – RTA + acetamide
2vc3 - RTA (mutant) + acetate
1il3, 1il4, 1il9 – RTA + guanine derivative
1ifu, 1fmp – RTA + formycin
1obt - RTA (mutant) + AMP
1apg – RTA + RNA
3px8 – RTA + formycin monophosphate
Ricin B chain (RTB)
3nbc, 3nbd – CnRTB + lactose – Clitocybe nebularis
3nbe – CnRTB + lactose derivative
3phz – RTB + glycoside – Polyporus squamosus
Ricin A+B chains
2aai - RTA + RTB
3rtj - RTA + RTB + dinucleotide
See Also
References
- ↑ 1.0 1.1 1.2 1.3 Lord JM, Roberts LM, Robertus JD. Ricin: structure, mode of action, and some current applications. FASEB J. 1994 Feb;8(2):201-8. PMID:8119491
- ↑ 2.0 2.1 2.2 Montfort W, Villafranca JE, Monzingo AF, Ernst SR, Katzin B, Rutenber E, Xuong NH, Hamlin R, Robertus JD. The three-dimensional structure of ricin at 2.8 A. J Biol Chem. 1987 Apr 15;262(11):5398-403. PMID:3558397
- ↑ Weston SA, Tucker AD, Thatcher DR, Derbyshire DJ, Pauptit RA. X-ray structure of recombinant ricin A-chain at 1.8 A resolution. J Mol Biol. 1994 Dec 9;244(4):410-22. PMID:7990130 doi:http://dx.doi.org/10.1006/jmbi.1994.1739
- ↑ Rutenber E, Ready M, Robertus JD. Structure and evolution of ricin B chain. Nature. 1987 Apr 9-15;326(6113):624-6. PMID:3561502 doi:http://dx.doi.org/10.1038/326624a0
- ↑ 5.0 5.1 Rapak A, Falnes PO, Olsnes S. Retrograde transport of mutant ricin to the endoplasmic reticulum with subsequent translocation to cytosol. Proc Natl Acad Sci U S A. 1997 Apr 15;94(8):3783-8. PMID:9108055
- ↑ Holmberg L, Nygard O. Depurination of A4256 in 28 S rRNA by the ribosome-inactivating proteins from barley and ricin results in different ribosome conformations. J Mol Biol. 1996 May 31;259(1):81-94. PMID:8648651 doi:10.1006/jmbi.1996.0303
- ↑ Iordanov MS, Pribnow D, Magun JL, Dinh TH, Pearson JA, Chen SL, Magun BE. Ribotoxic stress response: activation of the stress-activated protein kinase JNK1 by inhibitors of the peptidyl transferase reaction and by sequence-specific RNA damage to the alpha-sarcin/ricin loop in the 28S rRNA. Mol Cell Biol. 1997 Jun;17(6):3373-81. PMID:9154836
Contributions=
The following individuals contributed to the development of this page: Douglas Read, Michal Harel, Wayne Decatur, David Canner, Alexander Berchansky, Andrea Gorrell