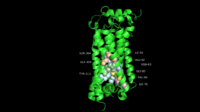
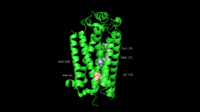
Dengue Virus Non-Structural Protein NS5
Introduction
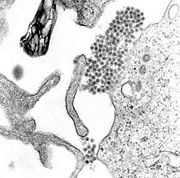
A TEM micrograph showing dengue virus virions (the cluster of dark dots near the center)
The dengue virus (DENV) belongs to genus Flavivirus, which also includes the West Nile Virus, Japanese Encephalitis Virus and Yellow Fever Virus. As a mosquito-borne pathogen, DENV causes dengue fever which can progress to dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS). Endemic to the tropics and subtropics, it is mainly transmitted by Aedes mosquitoes. Although there are 390 million dengue infections reported annually, no specific antiviral drug or vaccine has been yet developed [1].
Dengue fever manifests as a combination of severe headache, chills, backache and joint pain. DHF is a potentially deadly form of the fever characterized by sudden increase in body temperature and impaired blood homeostasis and clotting. DSS involves further impairment due to blood loss. Dengue virus can inhibit various mediators of the innate immune system such as interferons and bypass the inflammatory response[2].
There are four serotypes of the virus (DENV-1 – DENV-4) whose respective genomes share 60% sequence identity, with 90% sequence identity within a serotype[3]. The viral genome encodes three structural proteins (capsid, membrane and envelope) and seven non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5). By associating with host co-factors and each other, NS proteins form multi-protein replication complexes, which comprise the viral replication machinery. NS5 is also the most conserved among the viral proteins with at least 67% sequence similarity among the four serotypes and also shares some similarity with the NS5 protein of other flaviviruses[3][4].
Structure & Function
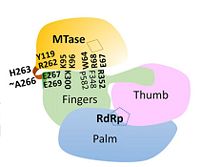
Overall structure of DENV NS5 highlighting functional domains and key inter-domain interactions
NS5 is a 104 kDa, 900 amino acids long protein, making it the largest of the DENV proteins. As per structural and biochemical studies, two functional domains exist within NS5. These being a S-adenosyl-L-methionine (SAM)-dependent methyltransferase (MTase) domain which spans residues 6-262 and a RNA-dependent RNA polymerase (RdRp) domain spanning residues 273-883 that are connected to each other by a flexible linker 10 amino acids in length (residues 263-272)[5].
Methyltransferase domain
The is involved in 5’ capping of the newly synthesized RNA - it catalyzes RNA cap methylation at both the N7 position on the guanosine and the 2’O on the subsequent nucleotide of the newly synthesized (+) RNA strand[4][5]. The MTase domain contains five parallel beta sheets and within NS5, lies above the finger subdomain of RdRp in such an orientation that its catalytic core (K61-D146-K180-E216) faces away from the inter-domain interface[5].
RNA-dependent RNA polymerase domain
The is responsible for the replication and transcription of the viral genome and new viral RNA is synthesized without the need for a primer (de novo RNA synthesis). Like other polymerases, the RdRp is similar in that it contains finger, thumb, and palm subdomains orientated in the canonical right-hand conformation as well as making use of a common catalytic mechanism for nucleotide incorporation involving two metal ions (zinc in NS5) located in the finger and thumb subdomains[3]. The entire RdRp domain is comprised of a total of 27 α-helices and seven β-sheets[3].
Together, the thumb and finger subdomains form a through which the RNA template and nucleotides enter en route to the catalytic core within the palm subdomain. Inside the tunnel, a priming loop protrudes from a thumb subdomain towards the active site where it controls access to and exit into the catalytic core and is thus critical in de novo initiation[3][4]. The RdRp is a feature unique to viruses and may be considered as a target for antiviral therapy.
Linker region
The defines the boundaries between the MTase and RdRp domains. It connects the two enzymatic domains acting like a swivel allowing the domains to adopt various orientations relative to each other. Residues 263-266, located after the MTase C-terminus, are key in this regard. They fold into a short 310 helix (though other conformations are also thought possible) resulting in compaction or elongation of the polypeptide chain and allowing for various inter-domain interactions over the course of the viral life cycle[5]. Apart from this flexibility, it has been demonstrated that the linker residues (268-272) N-terminal to the RdRp region enhance stability and polymerase activity as evidenced by results showing superior RdRp activity when the linker was present as compared to the RdRp alone, further corroborating that inter-domain interactions are required to successfully carry out RNA synthesis and viral replication and hence maintain infectivity[4][5][6].
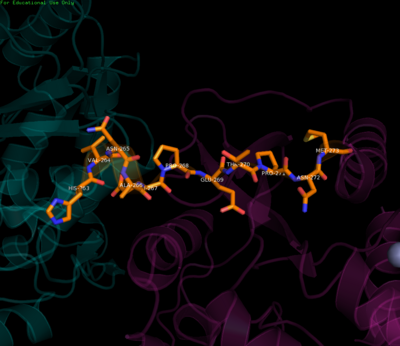
Residues of the linker region
Inter-domain interface
The majority of the stabilizing interactions within the inter-domain interface are polar in nature, resulting from contacts between charged side-chains as well as hydrogen-bonding mediated by trapped water molecules. Residues K95 and R352 form the major inter-domain polar contacts. The inter-domain is made up of two clusters with contributions from all three functional domains[5].
In the first cluster, interaction of linker residues with both RdRp and MTase domains is observed. Specifically, linker residue E267's side chain hydrogen bonds with residues Y119 and R262 of the MTase domain while E269 forms a salt-bridge with R361 of the RdRp domain. Polar interactions between residues from the MTase domain (K95 & K96) and the RdRp domain (E296-K300) also play an important role in the cluster.
The second cluster is based at residues (F348-K357) in the RdRp domain where guanidinium of R352 interacts electrostatically with a number of residues in the MTase domain (E67, E252, Q63). A salt-bridge is also formed between K357 and D256. Though hydrophobic interactions are not as numerous as hydrophilic interactions, they do exist as stacking interactions, namely between W64, R68, F348, and P582[5].
Through mutation studies in DENV-4, residues K95 and R353 have been specifically implicated in critical but non-enzymatic roles in virus RNA replication and infectivity while Y119 has been shown to be necessary for MTase activity and therefore a requirement for viral replication. Because of a majority of polar interactions as well as a small inter-domain buried surface area (1502 Å), it has been suggested that the two enzymatic domains may be able to associate and dissociate from each other with a relatively small energy penalty. This dynamic interaction may also lend itself to facilitating the recruitment of other viral and host proteins as part of the replication complex[5].
Conclusion
The inter-domain interface is characterized by a unique and electrostatic-rich network specific to DENV which enhances communication between and hence the function of the MTase and RdRp domains. Which, together with the linker region are essential in providing protein flexibility and therefore modulating viral replication and ultimately infectivity.