This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1656
From Proteopedia
| This Sandbox is Reserved from 26/11/2020, through 26/11/2021 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1643 through Sandbox Reserved 1664. |
To get started:
More help: Help:Editing |
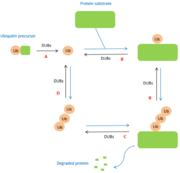
Deubiquitinases
| |||||||||||
References
- ↑ Mukhopadhyay D, Riezman H. Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science. 2007 Jan 12;315(5809):201-5. doi: 10.1126/science.1127085. PMID:17218518 doi:http://dx.doi.org/10.1126/science.1127085
- ↑ Schnell JD, Hicke L. Non-traditional functions of ubiquitin and ubiquitin-binding proteins. J Biol Chem. 2003 Sep 19;278(38):35857-60. doi: 10.1074/jbc.R300018200. Epub 2003, Jul 14. PMID:12860974 doi:http://dx.doi.org/10.1074/jbc.R300018200
- ↑ Amerik AY, Hochstrasser M. Mechanism and function of deubiquitinating enzymes. Biochim Biophys Acta. 2004 Nov 29;1695(1-3):189-207. doi:, 10.1016/j.bbamcr.2004.10.003. PMID:15571815 doi:http://dx.doi.org/10.1016/j.bbamcr.2004.10.003
- ↑ Urbe S, Liu H, Hayes SD, Heride C, Rigden DJ, Clague MJ. Systematic survey of deubiquitinase localization identifies USP21 as a regulator of centrosome- and microtubule-associated functions. Mol Biol Cell. 2012 Mar;23(6):1095-103. doi: 10.1091/mbc.E11-08-0668. Epub 2012, Feb 1. PMID:22298430 doi:http://dx.doi.org/10.1091/mbc.E11-08-0668
- ↑ Huang OW, Ma X, Yin J, Flinders J, Maurer T, Kayagaki N, Phung Q, Bosanac I, Arnott D, Dixit VM, Hymowitz SG, Starovasnik MA, Cochran AG. Phosphorylation-dependent activity of the deubiquitinase DUBA. Nat Struct Mol Biol. 2012 Jan 15;19(2):171-5. doi: 10.1038/nsmb.2206. PMID:22245969 doi:10.1038/nsmb.2206
- ↑ Huang OW, Ma X, Yin J, Flinders J, Maurer T, Kayagaki N, Phung Q, Bosanac I, Arnott D, Dixit VM, Hymowitz SG, Starovasnik MA, Cochran AG. Phosphorylation-dependent activity of the deubiquitinase DUBA. Nat Struct Mol Biol. 2012 Jan 15;19(2):171-5. doi: 10.1038/nsmb.2206. PMID:22245969 doi:10.1038/nsmb.2206
- ↑ https://authors.library.caltech.edu/261/1/AMBpb04.pdf
- ↑ Das C, Hoang QQ, Kreinbring CA, Luchansky SJ, Meray RK, Ray SS, Lansbury PT, Ringe D, Petsko GA. Structural basis for conformational plasticity of the Parkinson's disease-associated ubiquitin hydrolase UCH-L1. Proc Natl Acad Sci U S A. 2006 Mar 21;103(12):4675-80. Epub 2006 Mar 13. PMID:16537382
- ↑ Huang OW, Ma X, Yin J, Flinders J, Maurer T, Kayagaki N, Phung Q, Bosanac I, Arnott D, Dixit VM, Hymowitz SG, Starovasnik MA, Cochran AG. Phosphorylation-dependent activity of the deubiquitinase DUBA. Nat Struct Mol Biol. 2012 Jan 15;19(2):171-5. doi: 10.1038/nsmb.2206. PMID:22245969 doi:10.1038/nsmb.2206
- ↑ Alan D’Andrea, David Pellman,Deubiquitinating Enzymes: A New Class of Biological Regulators, Taylor and Francis Online, Sept 29 2008,DOI: https://doi.org/10.1080/10409239891204251
- ↑ Amerik AY, Hochstrasser M. Mechanism and function of deubiquitinating enzymes. Biochim Biophys Acta. 2004 Nov 29;1695(1-3):189-207. doi:, 10.1016/j.bbamcr.2004.10.003. PMID:15571815 doi:http://dx.doi.org/10.1016/j.bbamcr.2004.10.003
- ↑ Alan D’Andrea, David Pellman,Deubiquitinating Enzymes: A New Class of Biological Regulators, Taylor and Francis Online, Sept 29 2008,DOI: https://doi.org/10.1080/10409239891204251
- ↑ Singhal S, Taylor MC, Baker RT. Deubiquitylating enzymes and disease. BMC Biochem. 2008 Oct 21;9 Suppl 1:S3. doi: 10.1186/1471-2091-9-S1-S3. PMID:19007433 doi:http://dx.doi.org/10.1186/1471-2091-9-S1-S3
- ↑ Sun J, Shi X, Mamun MAA, Gao Y. The role of deubiquitinating enzymes in gastric cancer. Oncol Lett. 2020 Jan;19(1):30-44. doi: 10.3892/ol.2019.11062. Epub 2019 Nov 7. PMID:31897112 doi:http://dx.doi.org/10.3892/ol.2019.11062
- ↑ Saldana M, VanderVorst K, Berg AL, Lee H, Carraway KL. Otubain 1: a non-canonical deubiquitinase with an emerging role in cancer. Endocr Relat Cancer. 2019 Jan 1;26(1):R1-R14. doi: 10.1530/ERC-18-0264. PMID:30400005 doi:http://dx.doi.org/10.1530/ERC-18-0264
- ↑ David Komander, Michael J. Clague, Sylvie Urbé, Breaking the chains: structure and function of the deubiquitinases,Nature reviews molecular cell biology 10, August 2009, DOI:https://doi.org/10.1038/nrm2731