This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Bruna Oliveira de Almeida/Sandbox 1
From Proteopedia
SARS-CoV-2 RNA-dependent RNA polymerase
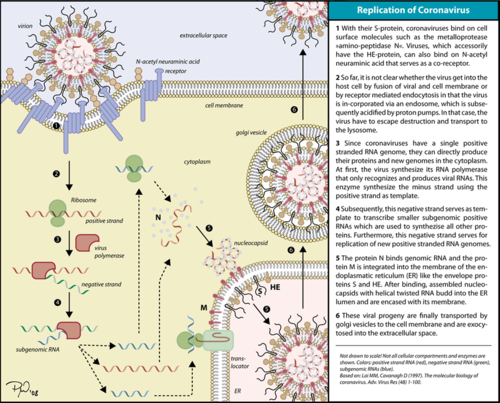
IntroductionThe RNA dependent RNA polymerase (RdRp) of SARS-CoV-2 (also known as nsp12), which is responsible for a major outbreak of the disease called Coronavirus Disease 2019 (COVID-19), declared pandemic by WHO in 11 march 2020, [1] is an enzyme that catalyzes the synthesis of RNA of the virus SARS-CoV-2.[2] This virus belongs to the betacoronavirus genus[3] and has a single stranded positive-sense RNA genome. Therefore, RNA dependent RNA polymerase plays a central role in the virus replication and livecycle. It has also been considered a good target for antiviral drugs.[2]
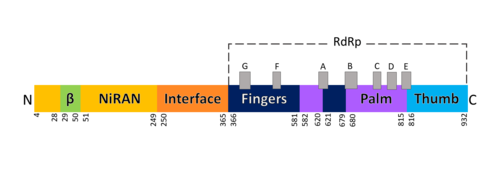
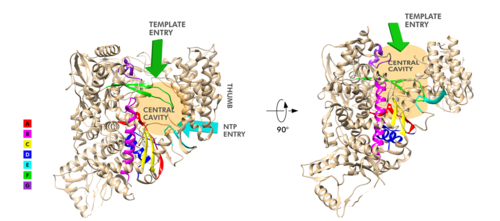
FunctionThe RdRp is responsible for replication and transcription of the virus genome alongside with other non-structural proteins (NSPs). After the firsts stages of infection, follow by the translation and assembly of the replicase complex, this enzyme helps the synthesis of genomic and sub-genomic RNA. Sub-genomic RNAs are used as mRNA for structural and accessory proteins.[4] The RdRp synthesizes negative-sense RNA and uses it as template for positive-sense strands for RNA replication for packaging into virus particles, and sub-genomic RNA transcription for translation.[5] Moreover, it has been reported that RdRp forms a complex with two other non-structural proteins, NSP7 and NSP8, which act as cofactors and confer processivity to its RNA-synthesizing activity.[6][7] StructureOverviewThe RdRp bonded to NSP7 and NSP8 consists of an approximately 160 kDa protein complex.[8] In fact, it was found that RdRp complexes with one NSP8 monomer and with one NSP7-NSP8 heterodimer, which help to stabilize the closed conformation of RdRp protein[2][9]. The RNA dependent RNA polymerase of SARS-CoV-2 contains 942 amino acid residues while NSP7 has 198 and NSP8 83 residues. Secondary structure is 38% (41 helices; 367 residues) and 13% (38 strands; 126 residues). To view the primary and secondary structure of SARS-CoV-2 RdRp and its cofactors visit rcsb [1]. The tertiary structure of RdRp protein of COVID-19 virus contains : a “right hand” (residues S367-F920), a (residues A4-T28 and T51-R249), which has a nidovirus RdRp-associated nucleotidyltransferase domain (NiRAN) architecture, an (residues A250-R365) that connects the RdRp domain and NiRAN domain, and a newly identified at its N terminus (residues D29-K50).[2] In the absence of DTT, it is found in the interface domain a between residues . RdRp domainThe RdRp domain has a conserved architectures and comprises : a (residues L366-A581 and K621-G679), a (residues T582-P620 and T680-Q815), and a (residues H816-E920).[2] In the absence of DTT, a is formed between residues , in the fingers subdomain.[2] Its active site consists of the polymerase A, B, C, D, E, F, and G, with important (759-SDD-761) been located in Motif C, (residues 753-FSMMILSDDAVVCFN-767), which is found in the palm subdomain, in the turn of .[2][9] (residues 611-TPHLMGWDYPKCDRAM-626) contains the classic divalent-cation-binding residue, conserved in most viral polymerases.[2]are close in the protein structure. As in other viral RNA polymerases, [10] the template-directed RNA synthesis is mediated by the RdRp domain motifs. The Motif G is established as a typical element of primer-dependent RdRp in some positive-sense RNA viruses also interacting with the primer strand to initiate RNA synthesis.[10] [2] The entry channel for the nucleoside triphosphate (NTP) is made up of a set of , as residues K545, R553 and R555 in Motif F, which help to stabilize the incoming nucleotide in the correct position for catalysis[2][9] by forming a fingertip that protrudes itself into the catalytic chamber, interacting with the finger extension loops and the thumb subdomain.[10] In general, most protein-RNA interactions involve the RNA phosphate-ribose back-bones and the double-stranded primer-template RNA is held by the fingers-palm-thumb subdomains.[9] NiRAN and β-hairpin domainThe complete SARS-CoV-2 RdRp protein structure obtained by cryo-EM[2][9] allowed to resolve the N-terminal portion of the as well to identify a N-terminal , that was in part unresolved for SARS-CoV RdRp protein.[8]The N-terminal portion of NiRAN can be seen (blue represents the N-terminus, red represents the C-terminus). Indeed, the resolution of residues 4 to 28 and 51 to 249 of NiRAN domain demonstrated that it comprises eight helices with a five-stranded β-sheet at the N-terminus.[2] It was also showed that residues form a β-strand that makes with the strand formed by residues , contributing to the stabilization of its conformation. Interestingly, in SARS-CoV RdRp these residues are less ordered. [2] Moreover, it was found that this β-hairpin is inserted in the groove clamped by the NiRAN domain and the palm subdomain and it forms close contacts that help to stabilize the overall structure.[2] NSP12-NSP7-NSP8 ComplexOn its own, the presents minimal polymerase activity, however, by bonding with NSP7 and NSP8 its polymerase activity is greatly stimulated.[11] Therefore, the is considered as the minimal core component for mediating SARS-CoV-2 RNA synthesis. Its structure highly resembles its counterpart from SARS-CoV. Although presenting some differences, these variations didn’t result in obvious structural changes.[10] This complex consists of one RdRp core catalytic subunit bound with two other structures that help its stabilization, an , and an ,[11][9][10] while the NSP8 monomer forms additional interactions with the interface domain and clamps the top region of the finger subdomain. The heterodimer bind above the thumb subdomain of NSP12 and clamps the finger extension loops in between stabilizing the adjacent fingertip loop. This interaction between the heterodimer and the RdRp is mainly mediated by the NSP7 portion of the heterodimer. And it’s interesting to notice that the two NSP8 subunits display different conformation consisting of substantial refold of the N-terminal extension helix region.[10] An attractive drug targetAs has been shown, SARS-CoV-2 RdRp protein is crucial for viral replication and so, it has been an important drug target in the fight against COVID-19.[2][12] In fact, growing attention has been given to RdRp as target of a class of antiviral drugs known as nucleotide analogs, including Remdesivir.[9][13] Remdesivir is an adenosine monophosphate analog and it is converted into its active drug form (its triphosphate form) within the cells.[14] It is positioned at the center of the catalytic active site and, as for other nucleotide analogs, Remdesivir was shown to inhibits the RdRp activity through non-obligate RNA chain termination, which requires the conversion of its monophosphate form to the triphosphate one.[9] In a recent study[9], it was described some differences between the apo and the complex (with Remdesivir and a template RNA) structures, showing some small conformational changes between them. (The three dimensional structure of the nsp12-nsp7-nsp8 complex bounded to the template-primer RNA and triphosphate form of Remdesivir can be seen in https://www.rcsb.org/3d-view/7BV2). References
| ||||||||||||