This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Carlos Henrique Passos/Sandbox1
From Proteopedia
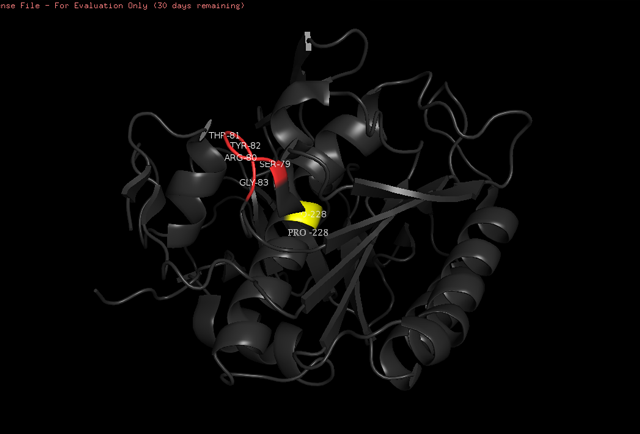
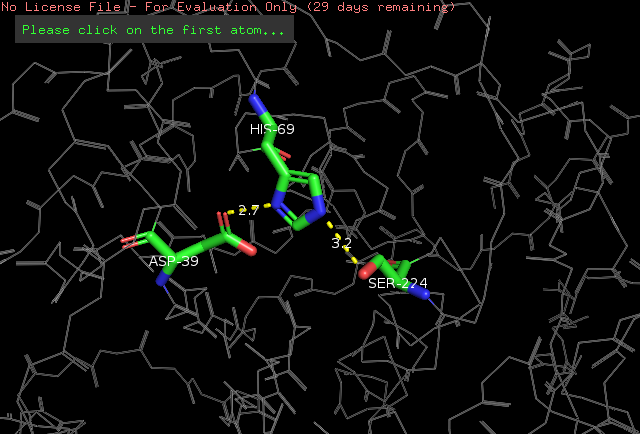
==Proteinase K==
| |||||||||||
References
- ↑ PDB 2ID8: Wang, Jiawei, Miroslawa Dauter, and Zbigniew Dauter. "What can be done with a good crystal and an accurate beamline?." Acta Crystallographica Section D: Biological Crystallography 62.12 (2006): 1475-1483. Available: https://www.ncbi.nlm.nih.gov/pubmed/17139083
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Cronier, Sabrina, et al. "Detection and characterization of proteinase K-sensitive disease-related prion protein with thermolysin." Biochemical Journal 416.2 (2008): 297-305. Available online: http://www.biochemj.org/content/416/2/297