This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Erin May/Sandbox 1
From Proteopedia
Contents |
Prions as a disease causing agent
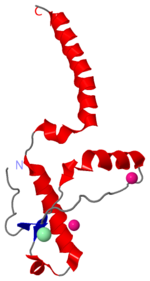
Prions are infectious or genetically coded misfolded proteins which act as templates upon which properly folded prion protein monomers can aggregate. Prions contain no nucleic acid such as other infectoius molecules or organisms. Human Prion Protein or Major Prion protein, exists as a normal constituent of human cells, found mostly in the brain[2] and is called PrPC.[3] PrPC is composed of mostly helix whereas the infectious form, PrPSc (also known as "scrapie" form), is composed of high percentage beta sheets.[3]
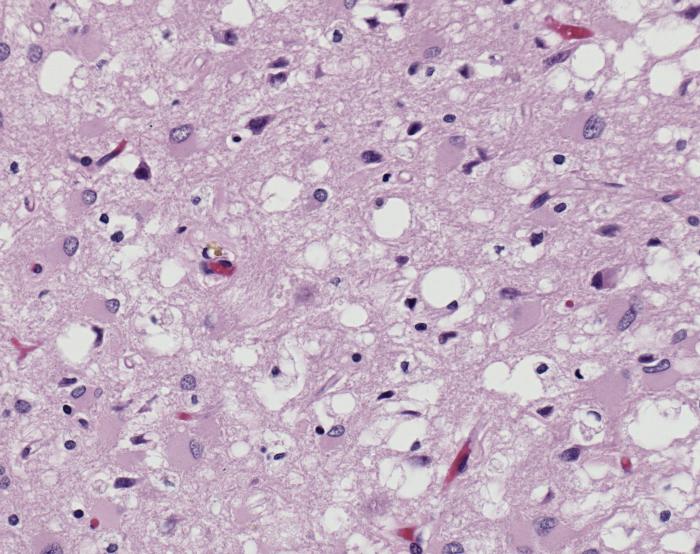
The diseases prions confer are neurodegenerative disorders which result from the large scale aggregation of these proteins. This "bubbles" of protein aggregates appear clear on a pictomicrograph and resemble a sponge. Bovine Spongiform Encephalopathy(BSE), or Mad Cow Disease, is a form of Transmissible Spongiform Encephalopathy caused by ingesting bovine prions. The first known cases of BSE occurred in the 1970's and have garnered a lot of media attention. Recently, feed bans in the United States and Canada have been adopted by the government in an attempt to stop the spread of BSE between cows. This bans the use of potential materials which would contain prion proteins, whether misfolded or wild-type. [4] For more information about the infections related to prions see Transmissible spongiform encephalopathy at Wikipedia.
Unfolding Mechanism
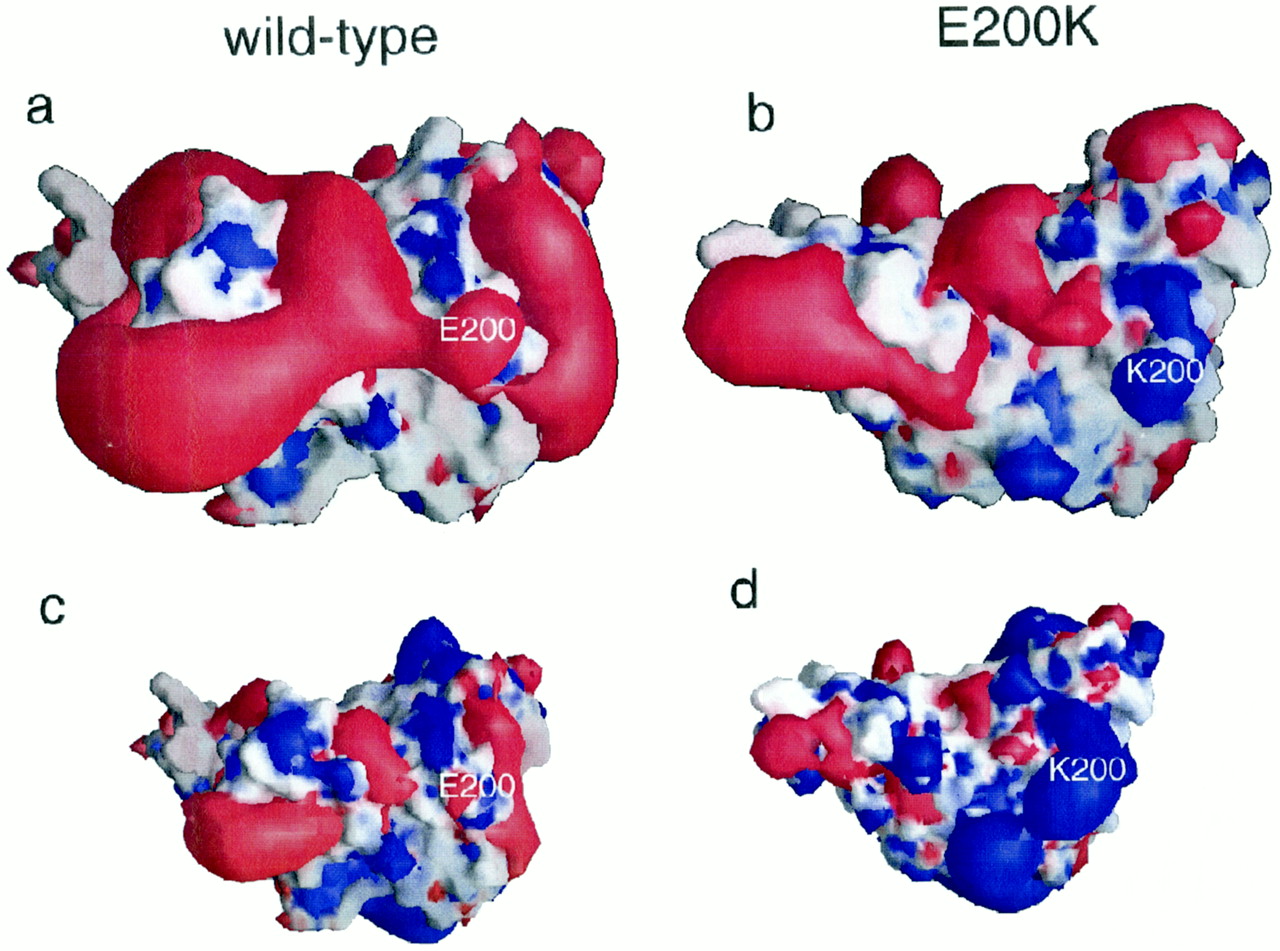
Currently, the mechanism by which a template prion unfolds a the helices of a properly folded prion protein is unknown. Specific residues have been shown to either confer resistance or lend themselves to this unfolding.
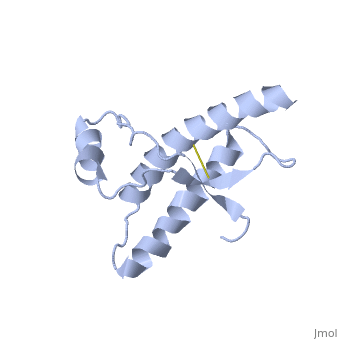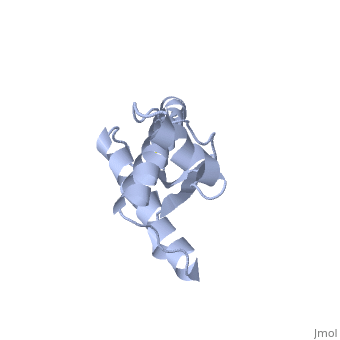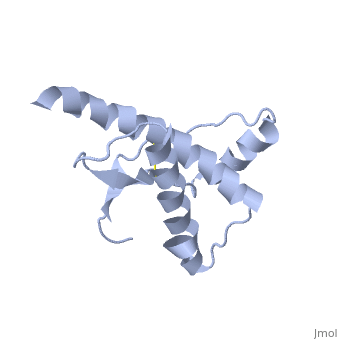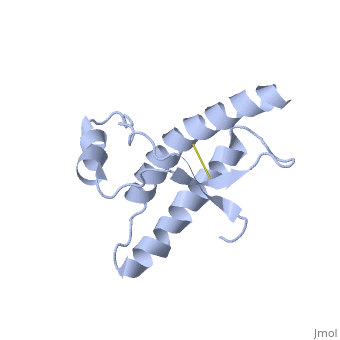
PrPC natural monomer
| |||||||||||
PrPSc
| |||||||||||
Dimer Form
| |||||||||||
Reference List
- ↑ Image of Creutzfeldt-Jakob positive brain tissue was obtained from The CDC's Public Health Image Library.
- ↑ Centers for Disease Control and Prevention: Prions. http://www.cdc.gov/ncidod/dvrd/prions/
- ↑ 3.0 3.1 Prusiner SB. Prions. Proc Natl Acad Sci U S A. 1998 Nov 10;95(23):13363-83. PMID:9811807
- ↑ Centers for Disease Control and Prevention: Bovine Spongiform Encephalopathy. http://www.cdc.gov/ncidod/dvrd/bse/index.htm
- ↑ 5.0 5.1 5.2 5.3 Lee S, Antony L, Hartmann R, Knaus KJ, Surewicz K, Surewicz WK, Yee VC. Conformational diversity in prion protein variants influences intermolecular beta-sheet formation. EMBO J. 2010 Jan 6;29(1):251-62. Epub 2009 Nov 19. PMID:19927125 doi:10.1038/emboj.2009.333
- ↑ 6.0 6.1 6.2 6.3 6.4 Knaus KJ, Morillas M, Swietnicki W, Malone M, Surewicz WK, Yee VC. Crystal structure of the human prion protein reveals a mechanism for oligomerization. Nat Struct Biol. 2001 Sep;8(9):770-4. PMID:11524679 doi:10.1038/nsb0901-770
- ↑ 7.0 7.1 7.2 7.3 Zhang Y, Swietnicki W, Zagorski MG, Surewicz WK, Sonnichsen FD. Solution structure of the E200K variant of human prion protein. Implications for the mechanism of pathogenesis in familial prion diseases. J Biol Chem. 2000 Oct 27;275(43):33650-4. PMID:10954699 doi:10.1074/jbc.C000483200