SerpinB5, also known as "mammary serine protease inhibitor" (), is considered a tumor suppressor serpin that does not present itself as a protein inhibitor like others of its own family, the serine protease inhibitor superfamily (serpins). Maspin was first identified in 1994 on mammary tissue and breast cancer cell lines [1], but it is also known to be expressed on a wide range of cell types and tissues, mainly in epithelial cells, i. e. in prostate, lung, skin, and corneal stromal cells [2]. It differs from ordinary serpins once it does not undergo the stressed (S) to relaxed (R) conformation which is a striking feature of other proteins in serpin’s superfamily [1]. Instead, its G-helix has quite a flexibility, capable of changing the conformation of the protein itself [3].
Maspin and its superfamily
Serpins
A serpin usually inhibit other proteins like serine proteases, caspases and papain-like cysteine proteases. However, some of them do not accomplish an inhibitory role. As an example, some of them function as hormone transporters, molecular chaperones or even as tumor suppressors [4]. Inhibitory serpins are considered “suicide molecules” because they can only be used once [5].
Serpins structure usually contain three ß-sheets (A, B and C) and eight to nine 𝛂-helices (hA-hI) on their structure, and the most important region to interact with their targets is the .The RCL is usually positioned out of the body of the serpins. When inhibiting proteases, serpins get their RCL cleaved out of the main structure, causing the amino-terminal portion of the RCL to form an additional fourth strand called s4A, once it is inserted into the center of ß-sheet A. This cleavage and modification on the structure of serpin is called the stressed (S) to relaxed (R) transition, in which the protein is in its biologically active state and transitions to a more thermal stable and latent state, respectively [4].
Maspin
Maspin is a 42 kDa protein [6] and is inserted in clade B of the serpin superfamily, composed of papain-like enzymes and inhibitory serpins that target cytotoxic apoptotic proteases which are working incorrectly [4]. Differently from other serpins, maspin does not undergo the S to R transition [4]. Instead, its is capable of undergoing a significant conformational change, that means this region of the molecule has some flexibility that allows movement. Accordingly, it is important to mention that studies have demonstrated, by superposing all of the maspin chains, a conformational heterogeneity at and around the G-helix (Fig. 1) [3].

Fig. 1 - Alignment and comparisons of maspin structures. A structural superposition of the 2.1 Å (PDB code
1wz9), a 2.8 Å (PDB code
1xu8) and a closed (PDB code
1xqg) maspin structures. Heterogeneity in and around the G-helix is shown in more details where G-helix from 2.1 Å structure is highlighted in
red, 2.8 Å structure in
yellow and closed maspin structure in
purple (INSPIRED BY doi:
http://dx.doi.org/10.1074/jbc.M412043200).
Also maspin is not limited to a certain cell compartment, once it is found on nucleus, cytoplasm, membrane, and as a secreted protein, according to the cell type and tissue [1][2]. Currently, it is known that the subcellular location of maspin is important for its tumor suppressor activity, and not only its protein levels inside the cell. In the past, there was a controversy about it, once maspin was upregulated in some tumors, while downregulated in others. Then, its translocation to the nucleus was observed and maspin’s nuclear localization was related to its tumor suppressor function [7]. However, contrary to what is expected, it has never been found a nuclear localization sequence (NLS), nuclear export sequence (NES), neither a secretory leader sequence (SLS) on maspin structure [8]. A map of maspin-like proteins is shown in Figure 2.
The tumor suppressor function of maspin is probably related to its activities, which are mainly inhibition of cell growth, invasion, tumoral migration, apoptosis stimuli, gene transcription regulation, angiogenesis inhibition [8] and prevention of oxidative damage of the proteome [9]. Besides all of these functions, maspin also has an important role in the organization of the epiblast during early embryonic development. However, maspin lacks studies on non-tumoral cell lines, and its role on a normal condition might be different from its activity inside a tumoral lineages [3].
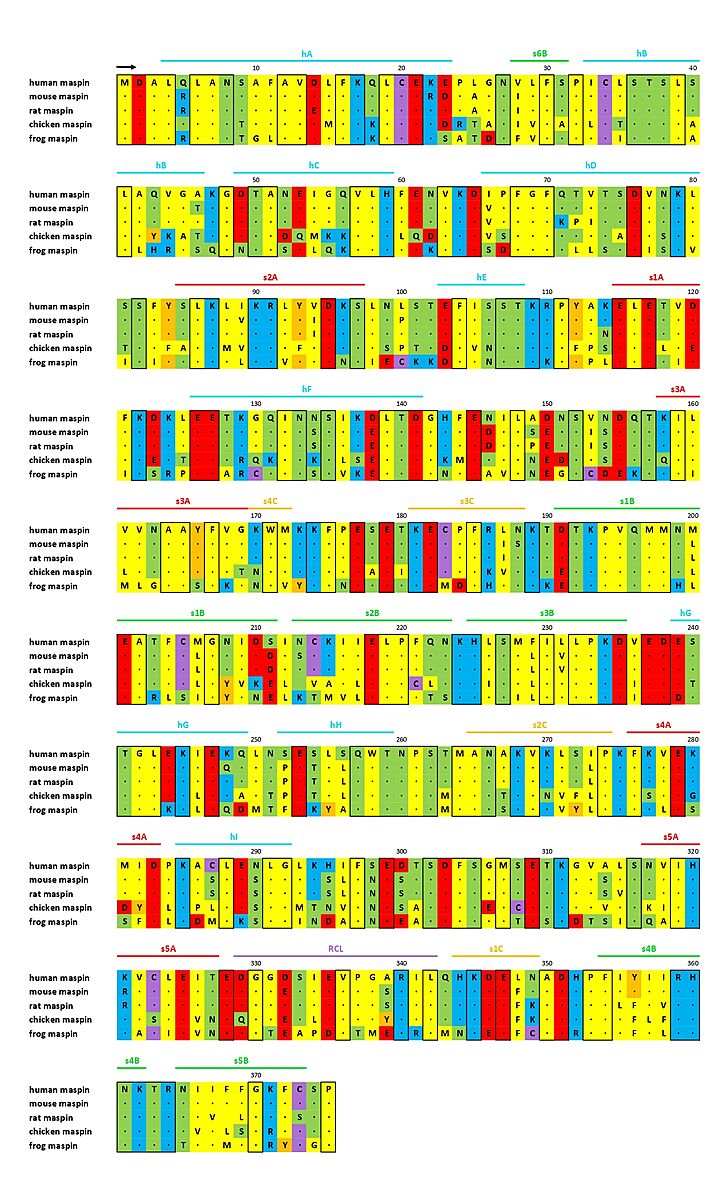
Fig. 2 - Sequence alignment of maspin-like proteins. The secondary structure of maspin is marked above the alignment and conserved residues are enclosed in a black box (ADAPTED FROM doi:
http://dx.doi.org/10.1074/jbc.M412043200).
Relevance
Maspin is a tumor suppressor molecule that has great potential to provide a prognosis for patients with tumor, since it seems to have a regulatory role in important functions mentioned above. In addition, it is thought that in the future, it could become a potential target for anti-tumor therapies to fight cancer cells [10].
Function and Structural highlights
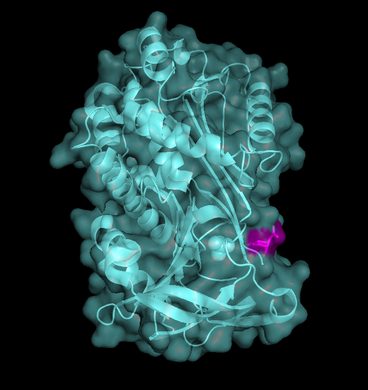
Fig. 3 - Maspin's ribbon/cartoon and surfice representation combined (PDB code
1wz9). RCL is highlighted in
pink.
RCL
does not differ a lot from other clade B serpins. It has three , nine and a (where you can find hydrophobic and polar residues in stick/wireframe representation, but also its and representations, this last one also combined with its ribbon/cartoon representation in Figure 3). The latter is also exposed in ordinary serpins and has a great flexibility.
Serpins that have mutations within their RCL which interfere with the ability to undergo the S to R conformational change cannot inhibit proteases and maspin’s RCL is the one among serpins that has the most different sequence [3][11][12].
Maspin does not present the conformational switch already discussed and does not have the consensus motif present in other serpins [3]. The intact RCL is necessary for maspin’s activity as a tumor suppressor [13], but there is no rearrangement of this structure, in other words, there is no S to R conformational change [14][15]. Besides that, the RCL alone has been related to cell matrix adhesion and inhibition of cell invasion [16].
One of the reasons for maspin’s RCL being unable to undergo the conformational switch is its limited flexibility, as it is not flexible like other serpins [3]. The RCL of maspin is shorter by four residues and lies closer to the core of the molecule, it is positioned further “back”, in other words closer to the N-terminal, than all of the other known serpin RCL structures. Besides that, the RCL of maspin is also stabilized by bonding interactions with (Fig. 4), leading to a more rigid structure. Additionally, the breach, which is presented in other serpins and where the cleaved RCL is inserted, is not seen on maspin [17].
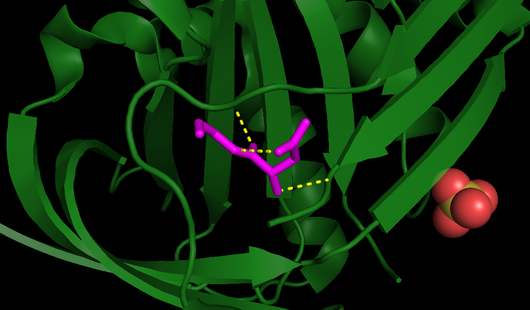
Fig. 4 - All RCL bonds, including those responsible for its stability (PDB code
1wz9). RCL is highlighted in
pink and all its bonds are displayed as
yellow dashed lines.
A curious phenomenon that happens with maspin is the aggregation of of tetramers, generating octamers in vitro. There is strong evidence that the hydrophobic residues on the RCL are responsible for the aggregation. The hydrophobic residues Val-336, Ile-341, Leu-342, Pro-337 and adjacent aminoacids present on the RCL are completely exposed to the solvent. Other types of intermolecular interactions, like minor salt links and hydrogen bonds between the s3C and s4C strands of opposing tetramers, also contribute to maintaining the structure, but the main force that results in the octamer are the hydrophobic associations [17]. Taking into account that the RCL is responsible for functions of cell matrix adhesion and inhibition of cell invasion [16], its hydrophobic nature is expected to be functionally important [17].
Further research is needed to understand if this phenomenon also happens in vivo conditions.
The RCL might be important for defining the protein subcellular localization. Modification of the Aspartate 346 (D346) by a glutamic acid (E) residue on the in maspin leaded maspin to a dominant nuclear distribution and increased interaction with HDAC1 in multiple cancer cell lines [18].
* Rainbow color schemes orientation:
Bulge around D and E-helices
Maspin also contains a buried between (where positive and negative charged, as well as neutral net charge are highlighted) inside a cavity formed by residues from , the B-helix and s2A. This intramolecular interaction causes a prominent at the N-terminal end of s1A and reveals a cavity below the D-helix. The salt bridge leads to a bulge around the D- and and is, possibly, a relevant cofactor recognition site. It is suggested that this salt bridge region, as in other serpins, may have some relevance at the interaction with a binding partner. Furthermore, the distortion in secondary structure caused by the salt bridge introduces into the center of a cluster of conserved positively charged waste. Once maspin is able to bind heparin, it is possible that these residues perform as a heparin binding site [3].

Fig. 5 - Maspin on its closed (PDB code
1xqg) and opened (PDB code
1wz9) form based on the ideia of the conformation "switch" in wich the rotation of G-helix changes maspin's charge distribuition
[3].
Positive and
negative charges can be seen in blue and red respectively, likewise
neutral net charge are represented by whyte color. The region comprising the G-helix is
highlighted.
G-helix
Maspin is able to undergo conformational change in and around the , presenting two shapes: an and a form. This is a real putative cofactor binding site and may determine maspin’s function.
Once the region at and around the G-helix has flexibility to undergo a conformational change, as a consequence, its (where positive and negative charges, as well as neutral net charge are highlighted) are reorganized and the central part of the helix structure becomes negative. The two structures, reflecting maspin in the open and closed conformation, show that rotation of the G-helix alters the local charge distribution (Fig. 5), suggesting that this movement represents a conformational “switch” (as we can see on these spacefill representations of and forms of maspin), what researchers have implied to be a cofactor binding site under conformational control based on G-helix negative charged patch modulation [3].
Physical interactions
It has been known that maspin interacts physically with a lot of different proteins. We will specifically discuss two of them below:
Histone Deacetylase 1 (HDAC1)
Maspin is able to bind to histone deacetylase 1 (HDAC1), which is up-regulated in many types of cancers and is an important class I nuclear deacetylase [18]. Either purified or endogenously expressed maspin is bound to and inhibits HDAC1. Maspin/HDAC1 interation in human protate tissues has been examined using GSH-affinity pull-down assay [19]. This interaction and, hence, the inhibition of HDAC1, may allow maspin to control a small set of genes involved in epithelial differentiation [20]. The inhibition of HDAC1 by maspin lead to increased acetylation of HDAC1 target protein Ku70, which in turn, caused an increase in apoptosis [21].
Unfortunately, it is not known where the exact binding site of maspin to HDAC1 is until the date of creation of this page (07/20/2022) and more studies are still needed.
Glutathione S-Transferase (GST)
The is an important molecule for the regulation of ROS-induced signaling and accomplishes an antioxidant role [22]. It is known that maspin interacts physically with GST, thus regulating the cell's response to oxidative stress [9].
Besides that, a new oxidized modification on maspin has been found, which has a different isoelectric point. The level of oxidized maspin increased in accordance with oxidative stress. Surprisingly, it was found that the oxidized form of maspin had lower binding affinity to GST. Researchers concluded that “the intramolecular disulfide-bonded formation of maspin might have some distinct properties compared to the native maspin isoforms under oxidative stress”.
Even though it was not possible to define exactly the molecular function of this new modification on maspin, it is presumed it might affect biochemical properties of the protein and its subcellular localization. Another hypothesis is that the modification might protect maspin from oxidative stress damages, which may result in an increase in the half-life of the protein [23].
Impressively, modification of the residue R340 of maspin RCL, to an Alanine, causes the protein to lose affinity with GST and leaded to a lower GST activity. Reseachers suspect that maspin might play a regulatory role instead of acting directly as a chaperone or as a detoxifying enzyme. Until the moment of creation of this webpage (07/20/2022), it has not been described the exact interaction site between GST and maspin, but researchers hypothesize an important role for the RCL on this interaction [9].





