From Proteopedia
proteopedia linkproteopedia link |
Introduction
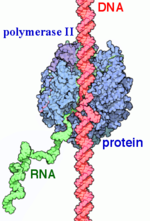 RNAP II transcription process. RNA Polymerases (RNAPs) are a group of enzymes that synthesize RNA in a process called transcription. During transcription the polymerase reads the DNA template strand and produces a RNA strand complementary to the template strand. The nascent RNA matches the DNA coding strand. Transcription can be divided into three processes that are discussed below: initiation, elongation and termination. Transcription in eukaryotes requires more distinct proteins for effective control. We see this as prokaryotic organisms have one core polymerase that synthesizes all of their RNA. However, eukaryotes have three distinct RNAPs named RNAP I, II, and III. RNAP I synthesizes rRNA precursors and RNAP III makes tRNA and the 5s rRNA. [1]
is responsible for the synthesis of pre-mRNA and snRNAs. It is 550 kDa and made of 12 (Rpb1-12) that range from 220-10 kDa. The subunits are highly conserved to the point that mammalian subunits can substitute with yeast subunits are there are little to no defects. [2] There are two large sub units and 10 smaller subunits, some of which are shared with RNAPs I and III. The active site of RNAP II is mostly made of the the two largest subunits Rpb1 and Rpb2.[1] The other subunits are involved in communication throughout the enzyme and help direct DNA and RNA into and out of the enzyme.
Rpb1 also has a that contains heptad repeats of YSPTSPS.[3] This region serves as the main control point for RNAP II. The CTD experiences various conformation changes based on it being in a hypo or hyper phosphorylation state. The heptad repeats may also exist in several forms with one repeat having 16 known states.[3] These states form a CTD code that is used to regulate the various stages of transcription and mRNA processing. The CTD is also shown to a part of cell cycle regulation as CDK/cyclins involved in the cell cycle modify the CTD and coordinate gene expression.[3]
While RNAP II is capable of transcription by itself it is non-selective of any particular DNA region. However some mutageneis studies have shown that RNAP II may have some role in selectivity.[1] To properly recognize regions upstream of the gene's transcription start site it requires several general transcription factors that are selective for these regions known as promoters and positions RNAP to accurately begin transcription.[2] There are other accessory proteins that regulate the rate of transcription based on environmental factors.[4]
History
RNA Polymerase was first discovered and isolated by Jerard Hurwitz in 1960. Prior to this, there was research in the synthesis of RNA. One enzyme known as polynucleotide phosphorylase was first isolated. It was initially thought to synthesize RNA but it was later discovered that it was DNA independent and later it was found to degrade RNA. This spurred Hurwitz to search for RNAP using E.coli extracts and in 1960 he showed reproducible RNA synthesis using his extracts and DNA. He published his findings along with three other labs who had also independently worked with RNAP. After this discovery, Hurwitz, along with John J. Furth, purified the enzyme from the E.coli extracts. The purified enzyme catalyzed RNA in the presence of rNTPs, DNA, and magnesium or manganese ions. [5]
Initially it was unknown if eukaryotes expressed one type of RNAP like eukaryotes or if there were multiple forms. In 1969 R. G. Roeder and and W. J. Rutter isolated three distinct species in sea urchin embryos by chromatography. They also showed that they required different environments for optimal activity and the forms are localized to different areas of the nucleus. RNAP I was found in the nucleous and RNAP II and III in the the nucleoplasm. Later experiments also showed that the different species if RNAP responded differently to to the inhibitor alpha-amanitin with RNAP being unresponsive to it, RNAP II inhibited by it, and RNAP II somewhere in between. Using this they could use varying concentrations of alpha-amanitin to see what types of RNA each RNAP was responsible for. [6]
Structural Components
This section will briefly discuss the chief structural components involved in the mechanism.
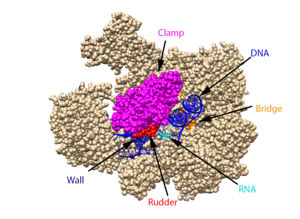 The clamp (magenta), wall (navy blue), rudder (red), bridge (orange), RNA (light blue), and DNA (blue) are depicted. See below for PDB's and residue numbers. To begin, the swings to trap the DNA in the cleft. Further along, the sends the DNA template through the cleft in approximately a 90° turn. Both the clamp and wall are parts of the Rpb2 subunit. Further along in the process, the separates the newly synthesized RNA strand from the DNA template. The DNA reforms into a double helix as it leaves RNA pol II.
Other components of RNA pol II include the following:
The jaw is the opening through which DNA enters. The funnel is what the NTP’s travel through to be incorporated into the growing RNA strand, and the pore is the end of the funnel. The is an Rpb1 segment that translocates the DNA-RNA complex at the end of each cycle of catalysis. is located within the active site and functions as the catalyst.
Transcription
Transcription can largely be divided into three sections: initiation, elongation, and termination. In the process of initiation, RNAP II recruits several general transcription factors (GTFs) to bind to the promoter region of the DNA, and this eventually forms the preinitiation complex (PIC). The DNA enters RNAP II through the clamp, and then it is unwound, creating a transcription bubble. With the DNA unwound, the active site of RNAP II catalyzes the synthesis of the first few RNA bonds. Once the carboxy-terminal domain (CTD) becomes phosphorylated, the clamp undergoes a conformation change to effectively trap the DNA, and a few of the GTFs dissociate, which changes complex to the Elongator complex.
After initiation, the process of elongation begins with the entry of NTPs. These NTPs largely enter through the funnel, and this tends to be a slow process because the funnel is only 12 Å in diameter, meaning that only one NTP can go through at a time. Once they have passed through the funnel, the NTPs enter which contains an Mg ion that is bound by three aspartate residues at positions D739, D741, and D743. If the NTP is complementary to the DNA strand, it is loaded onto the insertion site next to the RNA and held in place by the bridge. While in this insertion site, the metal ion and aspartate residues catalyze the reaction that forms a phosphodiester bond between the 3’ end of the RNA and the 5’ end of the NTP. This step then leads into a translocation step. To make room for the next NTP, there is a Brownian ratchet mechanism in which the nearby trigger loop undergoes a conformation change that causes the bridge to move to the next transition site. During the bridge’s transition, the DNA-RNA hybrid is partially help in place by the -amanitin. Once the bridge is in the new initiation site, the trigger loop returns to its original conformation, allowing the process to begin again.
In the termination stage, the CTD becomes dephosphorylated, which acts as a signal to dissociate the elongation complex. Once this signal is received, the new mRNA is released from RNP II through the rudder. Before being released, its 3’ end is polyadenylated. The DNA is brought back together at the other end of the transcription bubble, returning it to its original double-stranded form. Finally, RNAP II and the remaining GTFs dissociate from the DNA.
α-Amanitin
α-Amanitin is a bicyclic octapeptide that adheres tightly with RNAP II, which blocks the elongation steps. α-amanitin binds in the funnel and interacts with the bridge helix and adjacent Rpb1, but it does not inhibit the RNA pol II’s interaction with NTP. Instead, α-amanitin likely challenges the bridge’s conformational change that is necessary for the purposed RNAP translocation step. α-Amanitin, found in the poisonous mushroom death cap, leads to death after several days. This time frame aligns with the rate at which mRNA’s and proteins turnover.
 The chemical structure of α-amanitin.
General Transcription Factors
In both eukaryotes and prokaryotes, the basic mechanism for initiating transcription is the same: protein factors selectively bind to promoter regions on DNA. Prokaryotes use sigma factors while eukaryotes use a complex of 6 general initiation factors(GIFs)[4]. The combination of all the transcription factors bound to the DNA promoter region, in complex with RNAP II, is known as the preinitiation complex. This complex is necessary for an accurate initiation. The formation of the PIC occurs in an ordered pathway, beginning with the TATA box which is a promoter region on DNA at position -27.
PIC Formation
1. is highly conserved among eukaryotes. It recognizes and binds the TATA region of DNA. This is facilitated by a subunit named the TATA-binding protein (TBP) that has antiparralel beta-sheet that provides a large surface for minor groove interactions. This subunit binding also causes major deformations in the helix which may be important for further binding of the PIC units by creating a more compact protein-DNA complex. As the complex begins to form the TBP-TATA complex remains unchanged. Its shape resembles that of a saddle sitting on the DNA. [4]
2. > is the second to join the PIC. It is thought to be responsible for stabilizing the TBP/DNA complex and tethering the TFIID-DNA complex to RNAP I. It is also important in specifying the the TSS. In vitro studies have shown that accurate initiation can occur with only TFIID, TFIIB, and RNAP II suggesting that these two subunits serve to position RNAP. . Mutagenesis studies also suggest that it works as a spacer between TFIID and pol II. It may also function to ensure correct directionality. IF TBP binds the wrog end of TATA, TFIIB would have unfavorable interactions with TFB. [4]
3. binds directly to RNAP II and forms a very stable complex. It then escorts RNAP II to the promoter TFIIF also increases specificity and efficiency of transcription. It also acts similarly to bacterial sigma factor by inhibiting and reversing RNAP II binding to nonpromoter sites. TFIIF is a hetero-dimer of 30 and 70 kDa. [4]
4. is required to begin transcription even though RNAP II is bound to DNA before TFIIE binds. Once bound it recruits TFIIH. TfIIE is an α2β2 heterotetramer of 35 and 56 kDA. [4]
5. supports catalytic activity such as DNA ATPase, DNA helicase, and a kinase that phosphorylates the CTD of RPB1.[4](F) Some of its subunits are also components of DNA repair machinery. It is the last TF to bind and completes the .
6. is a co-activator that helps regulate PIC assembly. It was initially thought to be essential for activity. It binds to and stabilizes the early complexes. It also neutralizes transcription repressors. The mechanism is unknown but it is thought to either increase TBP affinity for DNA or displace repressors. TFIIA binds to the N-terminal of TBP. It lies upstream, of TATA where it can interact with promoter and enhancer elements.[4]
Once the is formed, initiates RNA synthesis and produces a short transcript. When RNAP II becomes phosphorylated, it releases some of the GTFs from the complex and moves away from the promoter. TFIID stays bound to the promoter and can reinitiate transcription and help with successive transcription. The transcription factors are replaced by a new six-protein complex call the Elongator. TFIIF and TFIIH both remain associated with RNAP II during elongation.
|
References
- ↑ 1.0 1.1 1.2 Young RA. RNA polymerase II. Annu Rev Biochem. 1991;60:689-715. doi: 10.1146/annurev.bi.60.070191.003353. PMID:1883205 doi:http://dx.doi.org/10.1146/annurev.bi.60.070191.003353
- ↑ 2.0 2.1 Myer VE, Young RA. RNA polymerase II holoenzymes and subcomplexes. J Biol Chem. 1998 Oct 23;273(43):27757-60. doi: 10.1074/jbc.273.43.27757. PMID:9774381 doi:http://dx.doi.org/10.1074/jbc.273.43.27757
- ↑ 3.0 3.1 3.2 Sobennikova MV, Shematorova EK, Shpakovskii GV. [C-terminal domain (CTD) of the subunit Rpb1 of nuclear RNA polymerase II and its role in the transcription cycle]. Mol Biol (Mosk). 2007 May-Jun;41(3):433-49. PMID:17685222
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 4.7 RNA polymerase II transcription initiation: A structural view
D. B. Nikolov, S. K. Burley Proceedings of the National Academy of Sciences Jan 1997, 94 (1) 15-22; DOI: 10.1073/pnas.94.1.15
- ↑ Hurwitz J. The discovery of RNA polymerase. J Biol Chem. 2005 Dec 30;280(52):42477-85. doi: 10.1074/jbc.X500006200. Epub 2005, Oct 17. PMID:16230341 doi:http://dx.doi.org/10.1074/jbc.X500006200
- ↑ doi: https://dx.doi.org/10.1038/nrm1796
(A)Young, Richard A. (2003-11-28). "RNA Polymerase II". Annual Review of Biochemistry. 60 (1): 689–715. doi:10.1146/annurev.bi.60.070191.003353. PMID 1883205.
(G) C-terminal domain of subunit Rpb1 of nuclear RNA polymerase II and its role in the transcription cycle
(B) https://www.jbc.org/content/273/43/27757 RNA Polymerase II Holoenzymes and Subcomplexes
(C) RNA polymerase II transcription initiation: A structural view
(D)The Discovery and Isolation of RNA Polymerase by Jerard HurwiMultiple forms of DNA-dependent RNA polymerase in eukaryotic organismstz
(E)
(e)https://www.nature.com/milestones/geneexpression/milestones/articles/milegene07.html
(F)The general transcription factors of RNA
polymerase II
(H)Interactions between the Human RNA Polymerase II Subunits*
Bushnell, D. A.; Westover, K. D.; Davis, R. E.; Kornberg, R. D. Structural Basis of Transcription: An RNA Polymerase II-TFIIB Cocrystal at 4.5 Angstroms. Science. 2004, 303, 983-988
Brueckner, F. and Cramer, P. Structural Basis of Transcription Inhibition by -amanitin and Implications for RNA Polymerase II Translocation. Nature Structure and Molecular Biology. 2008, 15, 811-818.
Cramer, P.; Bushnell, D. A.; Kornberg, R. D. Structural Basis of Transcription: RNA Polymerase II at 2.8 Ångstrom Resolution. Science. 2001, 292, 1863-1876
Evans, D. A.; Fitch, D. M.; Smith, T. E.; Cee, V. J. Application of Complex Aldol Reactions to the Total Synthesis of Phorboxazole B. J. Am. Chem. Soc. 2000, 122, 10033-10046.
Gnatt, A. L.; Cramer, P; Fu, J.; Bushnell, D. A.; and Kornberg, R. D. Structural Basis of Transcription: An RNA Polymerase II Elongation Complex at 3.3 Å Resolution. Science. 2001, 292, 1876-1882 1i6h
Hahn, S. Structure and Mechanism of the RNA Polymerase II Transcription Machinery. Nature Structure and Molecular Biology. 2004, 11, 394-403.
He, Yuan, et al. Near-atomic resolution visualization of human transcription promoter opening. Nature 533.7603. 2016.
Nudler, E. RNA Polymerase Active Center: The Molecular Engine of Transcription. Annu. Rev. Biochem. 2009, 78, 335-361.
Orphanides, George, Thierry Lagrange, and Danny Reinberg. The general transcription factors of RNA polymerase II. Genes & development 10.21. 1996. 2657-2683
Shah, N. et. al. Tyrosine-1 of RNA Polymerase II CTD Controls Global Termination of Gene Transcription in Mammals. Molecular Cell. 2018, 69, 48-61.
Uzman, A.; Voet, D. Student companion Fundamentals of biochemistry: life at the molecular level, 4th ed., Donald Voet, Judith G. Voet, Charlotte W. Pratt; John Wiley & amp; Sons, 2012.
Xu, J.; Lahiri, I.; Wang, W.; Wier, A.; Cianfrocco, M. A.; Chong, J.; Hare, A. A.; Dervan, P. B.; DiMaio, F.; Leschziner, A. E.; Wang, D. Structural Basis for the Initiation of Eukaryotic Transcription-coupled DNA Repair. Nature. 2017. 551, 653-657 5vvr
Xin, L.; Bushnell, D. A.; and Kornburg, R. D. RNA Polymerase II Transcription: Structure and Mechanism. Biochemica et Biophysica Acta. 2013, 1829, 2-8.
Yan, C., Dodd, T., He, Y., Tainer, J. A., Tsutakawa, S. E., & Ivanov, I. (2019). Transcription preinitiation complex structure and dynamics provide insight into genetic diseases. Nature Structural and Molecular Biology, 26(6), 397-406.
Alpha-aminitin chemical structure image courtesy of https://en.wikipedia.org/wiki/Alpha-Amanitin#/media/File:Alpha-amanitin_structure.png
Notes
From structural components:
Structural overview: [PDB: 5VVR: with highlighted sections mentioned below]
Bridge: Depicted: [PDB: 1I6H: 810-845.a]
Wall: Depicted: [PDB: 1R5U: 853-919.b; 933-972.b]
Clamp: Depicted: [PDB: 1R5U: 3-345.a; 1395-1435.a; 1158-1124.b]
Rudder: Depicted: [PDB: 5VVR: 306-321.a]
Content Donators
This page was created as a final project for the Advanced Biochemistry course at Wabash College during the Fall of 2019. This page was reviewed by Dr. Wally Novak of Wabash College.



