Keap1 is a BTB-Kelch substrate adaptor protein anchored to the actin cytoskeleton that regulates steady-state levels of Nrf2,[1] one of the main transcription factor of cytoprotective genes found in mammals.[2] Under homeostatic conditions, for ubiquitin-dependent degradation, repressing the expression of genes associated with cytological defense against highly reactive molecules, such as electrophilic chemicals, heavy metals and oxidative agents. [1] As such, when the cell is subjected to stresses of those kinds, Nrf2 is no longer targeted for ubiquitin-dependent degradation, thus allowing for the expressions of the genes responsible for the cellular defense against such reactive substances.
Relevant Keap1 information summary[3]
| Official Symbol
| KEAP1
|
| Official Full Name
| kelch like ECH associated protein 1
|
| Gene type
| protein coding
|
| RefSeq status
| REVIEWED
|
| Organism
| Homo sapiens
|
| Lineage
| Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Primates; Haplorrhini; Catarrhini; Hominidae; Homo
|
| Expression
| Ubiquitous expression in kidney (RPKM 10.4), prostate (RPKM 9.9) and 25 other tissues (see more)
|
| Orthologs
| Mouse, Cattle, Frog, and more (see more)
|
| Chromosomic Location:
| 19p13.2
|
Mechanism and function of Keap 1
In eukaryote cells, the defense against reactive molecules from both endogenous or exogenous sources are usually coordinated thanks to the expression of genes regulated by the Cap N' Collar transcription factors, a unique subset within the bZIP family of transcription factors.[4] Such molecules (reactive oxygen species, electrophilic chemicals and heavy metals), if not neutralized and eliminated, can affect the oxidation levels of most endocytic molecules and compromise the integrity of the DNA, leading to severe damage to the organism and causing many pathophysiological processes, including many known diseases in humans such as cancer[5]and cardiovascular problems. [6] The expression of genes regulated by the Cap N' Collar (CNC) set — which includes the transcription factors Nrf1, Nrf2, Nrf3, Bach1 and Bach2 — are indispensable for the induction of enzymes that can neutralize such reactive molecules, eliminate damaged macromolecules and restore cellular redox homeostasis.[1]
substrate adaptor protein responsible for regulating the levels of Nrf2, which together with Nrf1 constitutes the two most prevalent CNC transcription factors in regards to endocytic protection against reactive species in mammals, according to a 2003 experiment that consisted on the observation of the impact of individual CNC genes deletion.[7] Under basal conditions, Keap1 targets Nrf2 for ubiquitin-dependent degradation[1] by bridging the Neh2 domain (a conserved N-terminal regulatory domain) of the Nrf2 to a molecule of , a component of Cullin-RING E3 ubiquitin ligases complexes (CRLs). Once the lysine residues within the Neh2 domain of Nrf2 are marked, cyclical association and dissociation of this E3 ubiquitin ligase complex, mediated by the opposing actions of CAND1 and Cul3 neddylation, enables efficient ubiquitination of Nrf2[1], repressing the expression of Nrf2 dependent genes, reducing the ability of the cell to neutralize reactive molecules. But when the cell is under oxidative stress, a array of are modified,[8] perturbing the ubiquitin ligase complex and raising the levels of Nrf2, thus allowing for the expression of the genes responsible for the cytological defense against the invasive molecule. Although there are differences between the amino acid sequence of the Keap1 protein present in all eight organisms whose sequences have been stablished (mouse, rat, pig, human, Xenopus, zebrafish, Drosophila and rice)[9], studies have shown that in mice, the detection of xenobiotic agents is done by intermolecular disulfide bridges present in the Cys273 and Cys288 residues of the IVR domain, probably between C273 of one Keap1 molecule and C288 of a second[10].
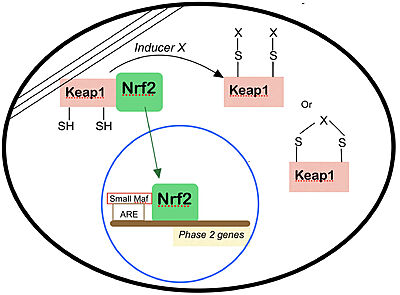
Nrf2 is anchored in the cytoplasm by binding to Keap1, which is attached to the actin cytoskeleton[8]. Inducers disrupt the Keap1–Nrf2 complex, and Nrf2 migrates to the nucleus where it forms heterodimers with other transcription factors such as small Maf that bind to the ARE regions of phase 2 genes and accelerate their transcription. Several types of modifications of Keap1 by inducers are shown.
Mechanism and function of Nrf2
Nrf2 belongs to the CNC (Cap ‘n’ Collar) family of b-Zip transcription factors, acting through the formation of a heterodimer with one of the small Maf proteins to activate the expression genes responsible for the defense against xenobiotic and oxidative stresses. [11] Like all CNC family members, Nrf2 is likely to interact with antioxidant responsive elements, manly: genes related to phase II enzymes production, genes related to the xc− system [12] (such as the xCT gene, responsible for encoding the cysteine–glutamate-exchange transporter, which mediates cysteine influx coupled with the efflux of intracellular glutamate) and the gene responsible for Mrp1 production (a ATP-binding cassette transporter that has an important role in the cellular exclusion of conjugated phase II metabolites[11]). Hence, Nrf2 coordinately regulates the xenobiotic conjugation reaction, the supply of intracellular glutathione and the excretion of xenobiotics, enabling efficient detoxification and cytoprotection against xenobiotic toxicity, and studies have shown that mice impaired with Nrf2 deficiency are more susceptible to xenobiotic stress. [13] [14] [15] [16]
According to Motohashi and Yamamoto,[11] Nrf2 is also an important regulator of oxidative-stress inducible genes, including heme oxygenase-1 and peroxiredoxin 1. In a 2002 study[17], a single-nucleotide polymorphism was detected in the promoter region of the Nrf2 gene of the mouse strain C57BL/6J, which is sensitive to hyperoxic stress. Supporting this, Nrf2-null mutant mice were found to be highly susceptible to hyperoxic lung injury. The impaired defense mechanisms against oxidative stress that are observed in the nrf2-null mutant mice could have resulted from the accumulation of reactive oxygen species (ROS) in the absence of Nrf2. A combination of electron paramagnetic resonance (EPR) and spin-probe kinetic analysis confirmed that there is a substantial decrease in the ability of nrf2-null mutant liver and kidney to eliminate ROS. This impaired elimination of ROS was exacerbated in aging female animals. Consistent with this result, old female Nrf2-deficient mice with an ICR genetic background often developed severe lupus-like autoimmune nephritis. Because ROS have a prominent role in the pathogenesis of nephritis, the accumulation of ROS due to Nrf2 deficiency must have exacerbated the mild glomerular lesions that are inherent to the ICR strain of mice.
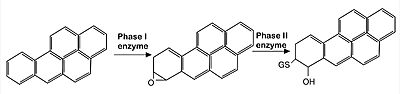
Image: representation of endocytic metabolisation of reactive agents (in this case, Benoz[a]pyrine molecules). The xenobiotic agent is first oxidized by phase I enzymes belonging to the P450 mono-oxygenase system (such as CYP1A1 and CYP1A2 enzymes). Since the product of such reactions are often highly reactive, they are subsequently detoxified by the induction of enzymes related to Phase II reactions (such as glutathione S-transferase and UDP–glucuronosyl-transferase), that promote the conjugation of phase I products with hydrophilic moieties, such as glutathione and glucuronic acid, so that the final, non-reactive product can be then eliminated by excretion. Nrf2 is the main CNC transcription factor responsible for the induction of those Phase II enzymes.
Thanks to the many binding sites of Nrf2, studies have shown that there are many alternative pathway from which Nrf2 degradation can occur, providing many mechanism for cellular control of Nrf2 activities [18], subjected to Keap1 regulation or otherwise. The following diagram from Warren L. Wu and Thales Papagiannakopoulos [19] summarizes all known Nrf2 related metabolic processes.
Structural and Genetic Information of Nrf2
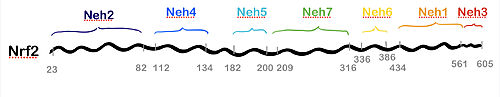
NRF2 is 605 amino acids long and possesses called NRF2-ECH homology (Neh) domains [20]. Their names and function is as follows:
(orange): A CNC-bZIP domain that allows Nrf2 to heterodimerize with small proteins.
(dark blue): allows for The Nrf2 peptide contains two short antiparallel β-strands connected by two overlapping type I β-turns stabilized by the aspartate and threonine residues. The β-turn region fits on the top face of the Kelch domain and the glutamate residues form multiple hydrogen bonds with highly conserved residues in Keap1.
, and domain (red, blue and light blue): may play a role in NRF2 protein stability and may act as a transactivation domain, interacting with component of the transcriptional apparatus (Neh3) or to a protein called cAMP Response Element Binding Protein (Neh4 and Neh5), which possesses intrinsic histone acetyltransferase activity.
(yellow): may contain a degron that is not recognized by Keap1, but it is instead involved in a redox-insensitive process of E3 ubiquitin ligase β-TrCP degradation of NRF2. This occurs even in stressed cells, which normally extend the half-life of NRF2 protein relative to unstressed conditions by suppressing other degradation pathways.
(green): interacts with the DNA-binding domain of retinoic X receptor alfa.
The gene responsible for Nrf2 coding in humans is called HAGRID 283, and its sequence of bases is available in the Human Ageing Genomic Resources database, and it is as follows:
Structural and Genetic Information of Keap1
Keap1 belongs to the metazoan superfamily of BTB-Kelch proteins, a widespread group of proteins that contain multiple Kelch motifs. The Kelch domain generally occurs as a set of five to seven kelch tandem repeats that form a . In , Keap1 has 624 amino acids, composing five different domains with the following names and caracteristics:
NTR: N-terminal region.
: The BTB/POZ domain (bric-a-brac, tramtrack, broad complex/Poxvirus zinc finger) is an evolutionary conserved domain also found in actin-binding proteins, zinc finger transcription factors, and substrate specific adaptor proteins in Cullin3 (Cul3)-based E3 ubiquitin ligase complexes [9]. In many cases, this is a protein-protein interaction domain that mediates dimerization, a process that is required for the binding with Nrf2.
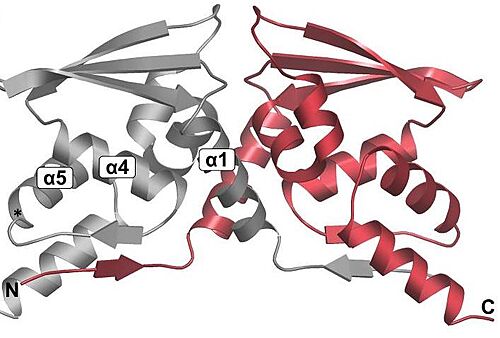
Overall fold of the Keap1 BTB crystallographic dimer as a cartoon representation. The N and C-termini, and key alpha-helical secondary structural elements are labelled for one BTB monomer.[21]. In mice, Keap1 dimmers are considered zinc metalloprotein, since the coupling is help by stoichiometric amounts of zinc and cobalt.
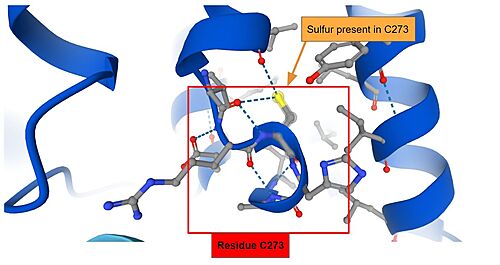
IVR: It is in the IVR domain that most of the cysteine residues is found, which suggest that this domain is the sensor for reactive/oxidative agents, since cysteine residues are highly reactive. As mentioned previously, it has been established that in mice, the residues Cys273 and Cys288 — both present in the IVR domain — contain the sulfite ions that are capable of reacting to invasive agents[9]. When the of C273 and C288 are perturbed by invasive substances, the Nrf2 that was once marked for ubiquitination is released and as such the cryoprotective reaction can occur. The sulfite present in Cys273 is represented bellow.
DGR (Kelch):The is a monomer and is comprised of six Kelch repeats that form a symmetric, six-bladed beta-propeller structure. The structure reveals that the Kelch repeat motif is defined by highly conserved glycine, tyrosine, and tryptophan residues. There are eight cysteine residues, none of them engaged in disulfide bonds. Interestingly, the first blade of the propeller consists of three strands from the N terminus of the protein and one strand from the C terminus, thus bringing the carboxy terminal region of Keap1 into close proximity to the IVR. Even thought the Kelch domain is not responsible for sensing the introduction of oxidative substances (that is the role of Cys273 and Cys288 in the IVR domain), it is still a highly reactive site and it is through the Kelch domain that the Keap1 protein is able to
CTR: C-terminal region. This region is of interest because it ends with a -CTC tripeptide, which is conserved in the other three mammalian proteins. This motif is very similar to the -CXXC- sequence found in the active site
of protein disulfide isomerase. Woycechowsky and Raines [22]have reported that the tripeptide CGCNH2 has a disulfide reduction potential similar to that of protein disulfide isomerase and even possesses disulfide isomerization activity. Whether the -CTC sequence in Keap1 has any disulfide reduction/isomerization activity has not yet been determined.
Structural highlights
Keap1 finalmente todos os dominios (humano):
Keap1 finalmente todos os dominios (mice):
Keap1 (mice) com cys 273 e 288 destacados:
BTB:
Kelch:
Nrf2 todos os dominios:
Nrf2 dom3:
Nrf2 dom1:
Nrf2 dom6:
Nrf2 dom7:
Nrf2 dom5:
Nrf2 dom4:
Nrf2 dom2:
Maf: