DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol [1] to the rescue.
Introduction
Histone background
HAT1 Background
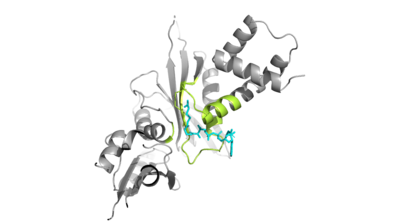
Hat1/Hat2 Complex Structure
Hat1 is not catalytically active until it binds with HAT2 to form the . HAT1 structure includes 317 residues and is identified as . HAT2 is identified as , which includes 401 residues. The activated complex acetylates residues in the 38 residue span of .
The HAT1 and HAT2 interface is stabilized by hydrogen bonds, salt bridges, and hydrophobic interactions. Most of these interactions are located in LP1 of the HAT1 domain, which forms a well-ordered helix. There are three major areas where hydrogen bonds are present. The side chain atoms of with the main chain nitrogen of Ala202 in HAT1. The side chain of makes hydrogen bonds with Leu288 and Phe205 respectively. The last area of hydrogen bonds between HAT1 and HAT is found between .
Mechanism
Of the five classes of HAT enzymes, the catalytic mechanisms for two of those enzymes, HAT1 and Rtt109, remains unclear. A structural overlay of HAT1 and Gcn5, a more well-known and understood HAT enzyme, found a conserved glutamate residue in the active site of both molecules. It was found that mutation at that active site glutamate residue greatly alters the catalytic ability of HAT1 and has been proven to be structurally important. [2] Using this information and structural information regarding the proximity of potentially catalytic residues, the most plausible mechanism for histone acetylation involves the following relevant residues and cofactors .
In this mechanism, the glutamate at residue 255 acts as a general base and deprotonates lysine 12 of histone 4 (the numbering of the modified lysine residue on histone 4 is shifted two residues).The deprotonated lysine then acts as a nucleophile and attacks the carbonyl carbon of acetyl coenzyme A (CoA, forming a tetrahedral intermediate transition state. The negative charge on the oxygen collapses down to for a double bond and the scissle bond between the carbonyl carbon and the sulfur atom of acetyl CoA is broken. The resulting product of this reaction is histone 4 with an acetyl-lysine at residue 12 and CoEnzyme A.
Application
References
Li, Y. et. al. Hat2p recognizes the histone H3 tail to specify the acetylation of the newly synthesized H3/H4 heterodimer by the Hat1p/Hat2p complex.(2014). Genes Dev.28:1217-1227. DOI:10.1101/gad.240531.114
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Li Y, Zhang L, Liu T, Chai C, Fang Q, Wu H, Agudelo Garcia PA, Han Z, Zong S, Yu Y, Zhang X, Parthun MR, Chai J, Xu RM, Yang M. Hat2p recognizes the histone H3 tail to specify the acetylation of the newly synthesized H3/H4 heterodimer by the Hat1p/Hat2p complex. Genes Dev. 2014 Jun 1;28(11):1217-27. doi: 10.1101/gad.240531.114. Epub 2014 May , 16. PMID:24835250 doi:http://dx.doi.org/10.1101/gad.240531.114
Student Contributors
- Caitlin Gaich
- Jordan Finch
- Morgan Buckley