Introduction
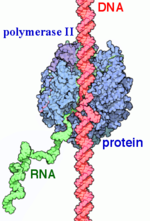
RNAP II transcription process.
RNA Polymerases (RNAPs) are a group of enzymes that synthesize RNA in a process called transcription. During transcription the polymerase reads the DNA template strand and produces a RNA strand complementary to the template strand. The nascent RNA matches the DNA coding strand. Transcription can be divided into three processes that are discussed below: initiation, elongation and termination. Transcription in eukaryotes requires more distinct proteins for effective control. We see this as prokaryotic organisms have one core polymerase that synthesizes all of their RNA. However, eukaryotes have three distinct RNAPs named RNAP I, II, and III. RNAP I synthesizes rRNA precursors and RNAP III makes tRNA and the 5s rRNA. [1]
is responsible for the synthesis of pre-mRNA and snRNAs. It is 550 kDa and made of 12 (Rpb1-12) that range from 220-10 kDa. The subunits are highly conserved to the point that mammalian subunits can substitute with yeast subunits are there are little to no defects. [2] There are two large sub units and 10 smaller subunits, some of which are shared with RNAPs I and III. The active site of RNAP II is mostly made of the the two largest subunits Rpb1 and Rpb2.[1] The other subunits are involved in communication throughout the enzyme and help direct DNA and RNA into and out of the enzyme.
Rpb1 also has a that contains heptad repeats of YSPTSPS.[3] This region serves as the main control point for RNAP II. The CTD experiences various conformation changes based on it being in a hypo or hyper phosphorylation state. The heptad repeats may also exist in several forms with one repeat having 16 known states.[3] These states form a CTD code that is used to regulate the various stages of transcription and mRNA processing. The CTD is also shown to a part of cell cycle regulation as CDK/cyclins involved in the cell cycle modify the CTD and coordinate gene expression.[3]
While RNAP II is capable of transcription by itself it is non-selective of any particular DNA region. However some mutageneis studies have shown that RNAP II may have some role in selectivity.[1] To properly recognize regions upstream of the gene's transcription start site it requires several general transcription factors that are selective for these regions known as promoters and positions RNAP to accurately begin transcription.[2] There are other accessory proteins that regulate the rate of transcription based on environmental factors.[4]
History
RNA Polymerase was first discovered and isolated by Jerard Hurwitz in 1960. Prior to this, there was research in the synthesis of RNA. One enzyme known as polynucleotide phosphorylase was first isolated. It was initially thought to synthesize RNA but it was later discovered that it was DNA independent and later it was found to degrade RNA. This spurred Hurwitz to search for RNAP using E.coli extracts and in 1960 he showed reproducible RNA synthesis using his extracts and DNA. He published his findings along with three other labs who had also independently worked with RNAP. After this discovery, Hurwitz, along with John J. Furth, purified the enzyme from the E.coli extracts. The purified enzyme catalyzed RNA in the presence of rNTPs, DNA, and magnesium or manganese ions. [5]
Initially it was unknown if eukaryotes expressed one type of RNAP like eukaryotes or if there were multiple forms. In 1969 R. G. Roeder and and W. J. Rutter isolated three distinct species in sea urchin embryos by chromatography. They also showed that they required different environments for optimal activity and the forms are localized to different areas of the nucleus. RNAP I was found in the nucleous and RNAP II and III in the the nucleoplasm. Later experiments also showed that the different species if RNAP responded differently to to the inhibitor alpha-amanitin with RNAP being unresponsive to it, RNAP II inhibited by it, and RNAP II somewhere in between. Using this they could use varying concentrations of alpha-amanitin to see what types of RNA each RNAP was responsible for. [6]
Structural Components
This section will briefly discuss the chief structural components involved in the mechanism.
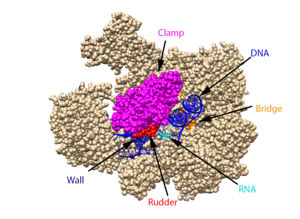
The clamp (magenta), wall (navy blue), rudder (red), bridge (orange), RNA (light blue), and DNA (blue) are depicted. See below for PDB's and residue numbers.
DNA enters through the jaw, an opening in RNAP II. The swings to trap the DNA in the cleft. The sends the DNA template strand through the cleft in a turn of approximately 90°. Both the clamp and wall are parts of the Rpb2 subunit. is located within the active site and functions as the catalyst. NTPs travel through the funnel to RNAP II’s active site, where they are incorporated into the growing RNA strand. The Trigger loop helps RNAP II select a substrate ribonucleotide by ensuring that only NTPs that form a Watson-Crick base pair with the DNA template base enter the active site of RNAP II. The phosphorylation of the C-terminal domain (CTD), which is a part of the Rpb1 subunit, triggers the conversion of RNAP II’s initiation complex to . The is an Rpb1 segment that translocates the DNA-RNA complex at the end of each catalytic cycle. Further along in the process, the separates the newly synthesized RNA strand from the DNA template strand, permitting the DNA double helix to reform as the DNA exits RNAP II.
Transcription
Pre-Initiation Complex
In both eukaryotes and prokaryotes, the basic mechanism for initiating transcription is the same: protein factors selectively bind to promoter regions on DNA [7][8]. Prokaryotes use sigma factors while eukaryotes use a complex of 6 general transcription factors (GTFs). These GTFs are all named similarly and begin with TF, for transcription factor, followed by the Roman numeral II since they are involved in transcription by RNAP II. The combination of all the transcription factors bound to the DNA promoter region, in complex with RNAP II, is called the pre-initiation complex (PIC)[7][8]. The formation of the PIC occurs in an ordered pathway, beginning with the upstream -35 promoter region, the -10 promoter region, and the transcription start site (TSS)[9] .
Process of PIC formation[7][10]:
1. is highly conserved among eukaryotes. It recognizes and binds the TATA region of DNA. This is facilitated by a subunit named the TATA-binding protein (TBP) that has antiparralel beta-sheet that provides a large surface for minor groove interactions. This subunit binding also causes major deformations in the helix which may be important for further binding of the PIC units by creating a more compact protein-DNA complex. As the complex begins to form the TBP-TATA complex remains unchanged. Its shape resembles that of a saddle sitting on the DNA. [4]
2. > is the second to join the PIC. It is thought to be responsible for stabilizing the TBP/DNA complex and tethering the TFIID-DNA complex to RNAP I. It is also important in specifying the the TSS. In vitro studies have shown that accurate initiation can occur with only TFIID, TFIIB, and RNAP II suggesting that these two subunits serve to position RNAP. . Mutagenesis studies also suggest that it works as a spacer between TFIID and pol II. It may also function to ensure correct directionality. IF TBP binds the wrog end of TATA, TFIIB would have unfavorable interactions with TFB. [4]
3. binds directly to RNAP II and forms a very stable complex. It then escorts RNAP II to the promoter TFIIF also increases specificity and efficiency of transcription. It also acts similarly to bacterial sigma factor by inhibiting and reversing RNAP II binding to nonpromoter sites. TFIIF is a hetero-dimer of 30 and 70 kDa. [4]
4. is required to begin transcription even though RNAP II is bound to DNA before TFIIE binds. Once bound it recruits TFIIH. TfIIE is an α2β2 heterotetramer of 35 and 56 kDA. [4]
5. supports catalytic activity such as DNA ATPase, DNA helicase, and a kinase that phosphorylates the CTD of RPB1.[4] Some of its subunits are also components of DNA repair machinery. It is the last TF to bind and completes the .
6. is a co-activator that helps regulate PIC assembly. It was initially thought to be essential for activity. It binds to and stabilizes the early complexes. It also neutralizes transcription repressors. The mechanism is unknown but it is thought to either increase TBP affinity for DNA or displace repressors. TFIIA binds to the N-terminal of TBP. It lies upstream, of TATA where it can interact with promoter and enhancer elements.[4]
Once the is formed, initiates RNA synthesis and produces a short transcript. When RNAP II becomes phosphorylated, it releases some of the GTFs from the complex and moves away from the promoter. TFIID, among other TFs, stays bound to the promoter region and can reinitiate transcription[11]. The transcription factors are replaced by a new six-protein complex called the Elongator. TFIIF and TFIIH both remain associated with RNAP II during elongation[11].
Initiation
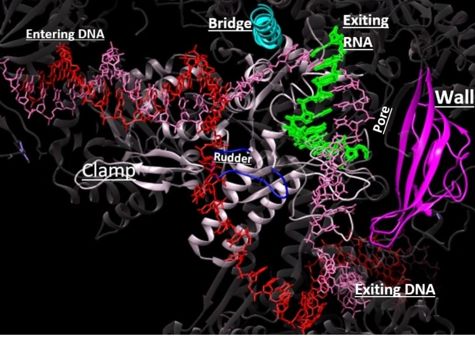
Depiction of RNAP II with DNA entering through the clamp (white), passing towards the wall (magenta). The rudder and bridge are shown in blue and teal, respectively. The growing RNA sequence is shown in green.
In the process of initiating transcription, RNAP II recruits several GTFs to bind to the promoter region of the DNA, and this eventually forms the PIC (described above)[7]. For this process to occur, the carboxy-terminal domain (CTD) must be unphosphorylated. It is unclear as to how the CTD and TFs interact, but it is known that the CTD must be in an unphosphorylated state for the TFs to form the PIC[12].
Once the proper GTFs are bound to the DNA and complexed with RNAP II, the DNA strands must be separated so an RNA transcript can be synthesized and read from the single stranded DNA (ssDNA) template. The DNA enters RNAP II through the clamp, and then 11-15 DNA bases at the TSS are separated through the “melting” of hydrogen bonds between bases, creating a in the DNA[11][13]. With the DNA strands unwound and in RNAP II, ssDNA passes through the of RNAP II. NTPs enter the complex through the pore, and bases are added to the growing RNA chain[11][14][15]. In the active site, magnesium ions help coordinate the incoming NTPs and form phosphodiester bonds[16]. These phosphodiester bonds form the backbone of the growing RNA segment.
In many systems this first sequence of several RNA bases (3-10) are termed abortive products because their purpose is to ensure RNAP II is correctly transcribing the RNA sequence before elongation occurs[11]. After roughly 30 RNA bases are synthesized, the CTD becomes phosphorylated in a transition to elongation[11][14]. CTD phosphorylation also releases some of the TFs complexed to DNA and RNAP II [12].
Reinitiation
Some of the TFs still remain on the DNA to reinitiate transcription and to mark where transcription has just occurred[11]. These remaining transcription factors are collectively called the . The scaffold complex consists of TFIIH, TFIIE, TFIID, and TFIIA and helps bypass the slow step of recruiting these TFs for reinitiation[11]. Thus, the scaffold complex only needs to recruit TFIIF, TFIIB, and RNAP II to restart transcription to synthesize another RNA product[11].
Elongation
After a short transcript is synthesized during initiation, the process of elongation is stimulated by a conformational transition[14]. To achieve the conformation conducive for elongation, the finger domain is displaced away from the active site to make room for the nascent RNA transcript and the CTD of the is phosphorylated[14][17]. Phosphorylation releases some initiation factors--some of these GTFs are left on the promoter region of the template to recruit another RNAPII[14]. has been thought to replace the GTFs on the phosphorylated CTD of Rpb1 to accelerate transcription through chromatin templates[14]. However, other recent studies have found evidence to the contrary and suggest a histone modification function of Elonagtor, acting through a chromatin- and acetyl-CoA-dependent mechanism [18].
DNA unwinds 3 nucleotides ahead of the active site (contained in Rpb1)[14]. Past the active site, the of Rpb2 redirects the template strand at about a 90 degree angle out of the cleft, orienting the in the active site towards the active site floor to align with the incoming NTP[14]. NTPs enter the active site on the floor side through a 12Å funnel called the pore, large enough for only one NTP at a time[14]. Once they have passed through the funnel, the NTPs enter the active site which is coordinated by three magnesium ions[19].
The domains of Rpb1 are found adjacent to the active site and have been implicated in RNAPII NTP selectivity, catalysis, and translocation[19][20]. The trigger loop oscillates between positions near downstream DNA and near the active site[19][20]. When a correctly paired NTP enters the A site, the trigger loop swings under the NTP, closing off the active site and allowing phosphodiester bond formation between the 3' end of the nascent RNA and the 5' end of the NTP[19]. The trigger loop is stabilized by extensive interactions with the bridge helix, the NTP ribose, base, and phosphate groups, and other active site residues[19][21]. The extensive trigger loop contacts slightly unwind and bend the bridge helix, a conformational change thought to play an important role in translocation[19]. of the trigger loop has been proposed to catalyze phosphodiester bond formation by acting as a general acid[19]. However, contradicting reports suggest that HIS1085 is not capable of acid catalysis, but rather catalyzes translocation as a positional catalyst[20]. Release of a pyrophosphate group after bond formation destabilizes interactions with His1085, releasing the trigger loop from the active site and allowing movement of the DNA-RNA hybrid helix and entrance of a new NTP[19][21][20]. Becasue the trigger loop maintains the bridge helix conformation, its release allows the bridge helix to relax and such movement likely contributes to RNAPII translocation[19].
Kinetics
Multiple kinetic models have been proposed with largely a consensus on some form of a Brownian ratchet mechanism involving the trigger loop and bridge helix in which the RNAPII has both forward and backward motion with a preference for forward translocation[22][20]. The net forward rate of RNAP II is about 2kb/min in vivo. RNAPII has nearly infinite processivity due to the Rpb2 subunit swinging down over DNA to trap it in the cleft[14].
Termination
The CTD in RNAP II's Rpb1 subunit has 26 heptapeptide repeats of the consensus sequence Tyr1-Ser2-Pro3-Thr4-Ser5-Pro6-Ser7[23]. The phosphorylation of Tyr1 results in the recruitment of elongation factors and helps prevent premature recruitment of termination factors during transcription[23]. The dephosphorylation of CTD ensures transition from the elongation stage to the termination stage. This dephosphorylation occurs at Tyr1 of the CTD and is catalyzed by the Glc7 subunit of the cleavage and polyadenylation factor (CPF)[23]. Dephosphorylated Tyr1 results in the recruitment of pcf11, a 3'-pre-mRNA processing factor. Pcf11 is able to bind to the phosphorylated Ser2 of CTD through its , resulting in the dissociation of RNAP II and the newly synthesized transcript from the DNA[23][24]. Before the transcript is released, its 3’ end is polyadenylated. The DNA is brought back together at the other end of the transcription bubble, returning it to its original double-stranded form[25].
α-Amanitin
is a bicyclic octapeptide that adheres tightly with RNAP II and inhibits elongation during transcription. The is beneath a bridge helix that extends between a region of Rbp1 (funnel) and a region of Rbp2[26]. The interacts with via a hydrogen bond and the interacts with residue Gln 768, which also interacts with residue His 816 of the bridge helix via a hydrogen bond[26]. There are also hydrogen bonding interactions between some residues on the funnel region of Rbp1 and α-amanitin[26]. The interaction between α-amanitin and the bridge helix restricts the movement of the bridge helix but it does not inhibit the entry of NTP into [27][26]. The movement of the bridge helix is required for the translocation process, and the binding of α-amanitin with the bridge helix likely prevents the bridge’s conformational change that is necessary for the purposed RNAP translocation step [27][26]. Inhibition of the translocation of DNA and RNA results in the active site not being emptied for the next round of synthesis [26]. α-Amanitin, found in the poisonous mushroom death cap, leads to death after several days. This time frame aligns with the rate at which mRNAs and proteins turnover[27].

The chemical structure of α-amanitin.




