use of JSmol in Proteopedia [1] article describing Jmol [2]
General Description
Human polymerase θ (pol θ) is large, 290kD enzyme consisting of three distinct domains [3][4]. An N-terminal helicase-like domain, whose exact cellular functions are a topic of on-going debate and research[5][6], is linked to a C-terminal, family A DNA polymerase domain by a large and disordered central region[4]. Notably, pol θ is the only known human polymerase to contain a polymerase and helicase domain in one molecule[7]. Crystal structures have been solved for the apo form of the helicase-like domain and the ternary complex of the polymerase domain. The focus of this wiki is the polymerase domain.
Cellular Functions
Pol θ is thought to promote overall genomic stability by performing several distinct cellular functions. The primary role of the enzyme is to repair of double-stranded DNA breaks as the key enzyme in an error-prone, non-homologous end-joining pathway called alternative end-joining[8] or theta-mediated end-joining[9]. Other functions include translesion synthesis, the ability of the polymerase to bypass and extend past a site of oxidative DNA damage[10], base excision repair [11], and possibly DNA replication timing [12]. Pol θ has the specialized ability to extend DNA from minimally-paired primers[13]. Repair by this enzyme is considered to error-prone due to its tendency to add or delete short indels [8].
Disease
Several types of cancer, such as breast, ovarian, and oral carcinomas, have shown significantly higher expression levels of pol θ and correlate to poorer patient outcomes[14][15][16]. Genomic studies have shown that more than half of epithelial ovarian cancers have defects in the error-free repair pathway of homologous recombination[17] and, as a result, have an increased dependence on theta-mediated end-joining [14]. Double-stranded break repair by pol θ may be thought of as a "backup" pathway which cells depend on more when the machinery involved in homologous recombination is compromised or otherwise unavailable. This enzyme has been identified as a potential therapeutic target due to overexpression in cancers in combination with studies that have shown inhibition of pol θ to sensitize human and mouse cells to radiation and chemical agents which induce double-stranded breaks[14][18][19].
Structural Highlights
Two crystal structures of the polymerase domain have been solved bound to DNA to a resolution of 3.9Å; inserting ddATP opposite tetrahydrofuran (THF, representing an abasic site) with 4 molecules in the asymmetric unit and inserting ddGTP opposite dCMP with 2 molecules in the asymmetric unit[7]. An overall assessment of the structures display the canonical . The DNA is thought to "sit" on the palm and is enclosed by the thumb and fingers via the minor groove. The palm houses the active site and the fingers move from an open, unbound position to a closed position when bound to DNA, promoting synthesis. The closed conformation of the polymerase domain becomes obvious when compared to the , such as Thermus aquaticus DNA polymerase I.
ddATP Opposite THF
An inspection of the active site reveals the , in this instance calcium. This structure required the use of calcium, a known inhibitor of polymerase activity, as the primer strand retains the 3' hydroxyl which would otherwise be subjected to nucleophilic attack. The similarly conserved .
dCMP Opposite ddGTP
from the same study was solved with magnesium as the coordinated metal, a 27 amino acid N-terminal truncation, and a blunted DNA oligomer to remove the 3' template overhang. The overall structure is virtually identical to the aforementioned structure complexed with calcium, with the exception of a slight shift in position of the O-helix to be closer to the cognate C:G basepair.
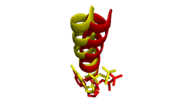
Alignment of O-helices and incoming nucleotides. Mg2+ complex (yellow) shifts slightly closer to incoming ddGTP than Ca2+ complex (red) relative to its incoming ddATP.
Structural Insights into Function
Proofreading Activity
Family A DNA polymerases harbor an N-terminal exonuclease domain and generally conserve the overall fold. Some members of this family, e.g. E. coli pol I, display active 3'-5' proofreading activity via . Other members, such as pol θ, do not harbor proofreading activity[7] and these three coordinating residues are notably absent at structurally equivalent positions (see below).

Exonuclease domains of active pol I (left) and inactive pol θ (right). Three coordinating residues and metal are absent in pol θ.
Translesion Synthesis
An R2254V variant was made to investigate the importance of this residue in pol θ's ability to extend single-stranded DNA and bypass abasic sites and bulky thymine glycol lesions.
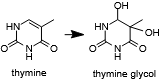
Thymine may oxidized to form a lesion that must be repaired or bypassed.
Bacterial family A polymerases which do not have this ability retain a valine or leucine at the equivalent position. The R2254V mutant retained its ability to extend double-stranded but not single-stranded DNA and also was not able to bypass abasic sites or thymine glycol. These findings indicated to the authors that is required to compensate for the missing contacts to the template strand due to a lesion
[7].
Sticky Thumb
In addition to R2254, additional upstream contacts to the primer DNA strand mediated by and of the thumb subdomain can be observed. These three salt bridges provide specialized contacts that are not observed in other family A polymerases, in addition to R2201 and R2315 which are also present in pol ν and pol I. Alanine substitutions of K2181, R2202, and R2254 resulted in inhibition of pol θ's ability to bypass an abasic site or thymine glycol. These findings prompted the authors to assert that this may be responsible for the heightened ability of pol θ to synthesize across lesions and to extend from minimally-paired primers[7].
Evolutionarily Related Proteins
DNA polymerases are surprisingly diverse structurally when compared to other enzymes[20].Polymerase θ is a family A (aka pol I) DNA polymerase, based on shared sequence and structural characteristics. The structure of the palm domain and the ability to coordinate 2 divalent metal ions to catalyze nucleotidyl transfer is shared among all DNA polymerases, with the exception of the pol β family[20][21]. The major differences are located in the thumb, fingers, and exonuclease subdomains. Some examples include that Family A polymerases have additional, conserved interactions with the DNA backbone in the thumb domain and also retain the relative location of the exonuclease subdomain[20]. It is important to note that major differences can exist even within a single family, such as an active or inactive exonuclease subdomain, as discussed previously in this article.
Links to other family A DNA polymerases:
T7 DNA polymerase
Geobacillus stearothermophilus DNA polymerase I
E. coli polymerase I
Thermus aquaticus DNA polymerase
Human polymerase γ (subunit 2)




