This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2vab
From Proteopedia
(Difference between revisions)
| (13 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | [[Image:2vab.gif|left|200px]] | ||
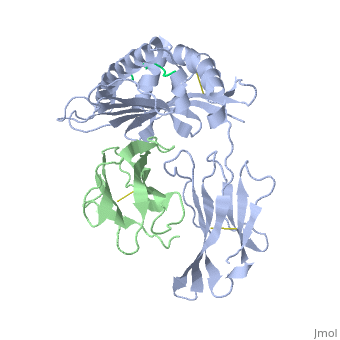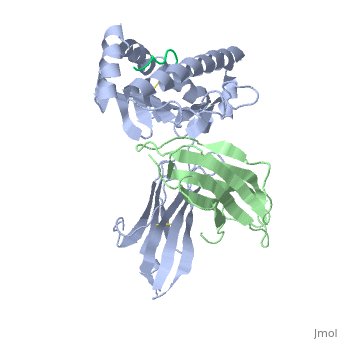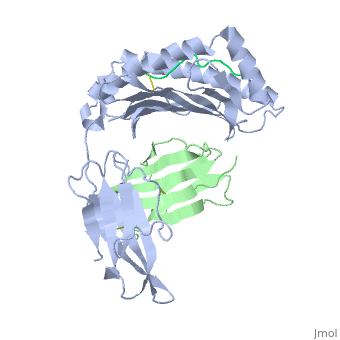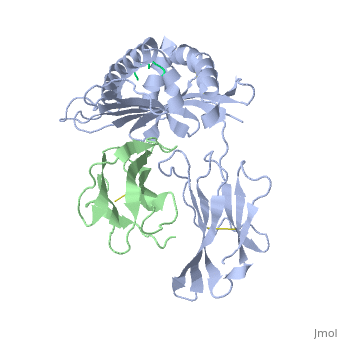
| - | + | ==MHC CLASS I H-2KB HEAVY CHAIN COMPLEXED WITH BETA-2 MICROGLOBULIN AND SENDAI VIRUS NUCLEOPROTEIN== | |
| - | + | <StructureSection load='2vab' size='340' side='right'caption='[[2vab]], [[Resolution|resolution]] 2.50Å' scene=''> | |
| - | + | == Structural highlights == | |
| - | + | <table><tr><td colspan='2'>[[2vab]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/Lk3_transgenic_mice Lk3 transgenic mice] and [https://en.wikipedia.org/wiki/Murine_respirovirus Murine respirovirus]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1vab 1vab]. The February 2005 RCSB PDB [https://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''Major Histocompatibility Complex'' by David S. Goodsell is [https://dx.doi.org/10.2210/rcsb_pdb/mom_2005_2 10.2210/rcsb_pdb/mom_2005_2]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2VAB OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2VAB FirstGlance]. <br> | |
| - | + | </td></tr><tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">BETA-2-MICROGLOBULIN ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10090 LK3 transgenic mice])</td></tr> | |
| - | | | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2vab FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2vab OCA], [https://pdbe.org/2vab PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2vab RCSB], [https://www.ebi.ac.uk/pdbsum/2vab PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2vab ProSAT]</span></td></tr> |
| - | + | </table> | |
| - | + | == Function == | |
| - | + | [[https://www.uniprot.org/uniprot/HA1B_MOUSE HA1B_MOUSE]] Involved in the presentation of foreign antigens to the immune system. [[https://www.uniprot.org/uniprot/NCAP_SENDE NCAP_SENDE]] Encapsidates the genome in a ratio of one N per six ribonucleotides, protecting it from nucleases. The nucleocapsid (NC) has a helical structure with 13.07 N per turn. The encapsidated genomic RNA is termed the NC and serves as template for transcription and replication. Replication is dependent on intracellular concentration of newly synthesized N, termed N(0), which corresponds to the protein not associated with RNA. In contrast, when associated with RNA, it is termed N. During replication, encapsidation by N(0) is coupled to RNA synthesis and all replicative products are resistant to nucleases. [[https://www.uniprot.org/uniprot/B2MG_MOUSE B2MG_MOUSE]] Component of the class I major histocompatibility complex (MHC). Involved in the presentation of peptide antigens to the immune system. | |
| - | + | == Evolutionary Conservation == | |
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/va/2vab_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=2vab ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | The x-ray structures of a murine MHC class I molecule (H-2Kb) were determined in complex with two different viral peptides, derived from the vesicular stomatitis virus nucleoprotein (52-59), VSV-8, and the Sendai virus nucleoprotein (324-332), SEV-9. The H-2Kb complexes were refined at 2.3 A for VSV-8 and 2.5 A for SEV-9. The structure of H-2Kb exhibits a high degree of similarity with human HLA class I, although the individual domains can have slightly altered dispositions. Both peptides bind in extended conformations with most of their surfaces buried in the H-2Kb binding groove. The nonamer peptide maintains the same amino- and carboxyl-terminal interactions as the octamer primarily by the insertion of a bulge in the center of an otherwise beta conformation. Most of the specific interactions are between side-chain atoms of H-2Kb and main-chain atoms of peptide. This binding scheme accounts in large part for the enormous diversity of peptide sequences that bind with high affinity to class I molecules. Small but significant conformational changes in H-2Kb are associated with peptide binding, and these synergistic movements may be an integral part of the T cell receptor recognition process. | ||
| - | + | Crystal structures of two viral peptides in complex with murine MHC class I H-2Kb.,Fremont DH, Matsumura M, Stura EA, Peterson PA, Wilson IA Science. 1992 Aug 14;257(5072):919-27. PMID:1323877<ref>PMID:1323877</ref> | |
| + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| + | </div> | ||
| + | <div class="pdbe-citations 2vab" style="background-color:#fffaf0;"></div> | ||
| - | == | + | ==See Also== |
| - | + | *[[Beta-2 microglobulin 3D structures|Beta-2 microglobulin 3D structures]] | |
| - | + | *[[Cation-pi interactions|Cation-pi interactions]] | |
| - | == | + | *[[MHC 3D structures|MHC 3D structures]] |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| + | [[Category: Large Structures]] | ||
| + | [[Category: Lk3 transgenic mice]] | ||
[[Category: Major Histocompatibility Complex]] | [[Category: Major Histocompatibility Complex]] | ||
| - | [[Category: | + | [[Category: Murine respirovirus]] |
| - | [[Category: | + | [[Category: RCSB PDB Molecule of the Month]] |
| - | + | [[Category: Fremont, D H]] | |
| - | [[Category: Fremont, D H | + | [[Category: Wilson, I A]] |
| - | [[Category: Wilson, I A | + | [[Category: Class i major histocompatibility complex]] |
| - | [[Category: | + | [[Category: Histocompatibility antigen]] |
| - | [[Category: | + | [[Category: Mhc i]] |
| - | [[Category: | + | [[Category: Mhc-peptide complex]] |
| - | [[Category: | + | |
| - | + | ||
| - | + | ||
Current revision
MHC CLASS I H-2KB HEAVY CHAIN COMPLEXED WITH BETA-2 MICROGLOBULIN AND SENDAI VIRUS NUCLEOPROTEIN
| |||||||||||