This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1694
From Proteopedia
(Difference between revisions)
| Line 25: | Line 25: | ||
== Other important features == | == Other important features == | ||
| + | |||
In the structure INPP1D54A, there is a mutation in the amino acid aspartic acid (D)54 and causes it to change to alanine (A)54 (<scene name='89/892737/Alanine/1'>mutation in D54</scene>). This mutation does not impact the substrate affinity but does decrease the activity of INPP1. <ref name="dollins" /> | In the structure INPP1D54A, there is a mutation in the amino acid aspartic acid (D)54 and causes it to change to alanine (A)54 (<scene name='89/892737/Alanine/1'>mutation in D54</scene>). This mutation does not impact the substrate affinity but does decrease the activity of INPP1. <ref name="dollins" /> | ||
[[Image:Motif.png | thumb]] | [[Image:Motif.png | thumb]] | ||
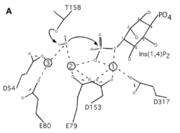
| - | + | A six amino acid <scene name='89/892737/Motif/1'>motif</scene>, DPIDxT anchors the metal-binding sites in the protein that are likely involved in catalysis while the metal binds to the substrate. <ref name="dollins" /> The sixth amino acid, x, is not as important as the other five, however, it can be any amino acid depending on the related crystallized structure to INNP1D54A or to a similar protein. | |
Lithium is an uncompetitive inhibitor for this protein and when it binds to a metal site it causes the protein not to function. | Lithium is an uncompetitive inhibitor for this protein and when it binds to a metal site it causes the protein not to function. | ||
Revision as of 02:56, 9 December 2021
| This Sandbox is Reserved from 10/01/2021 through 01/01//2022 for use in Biochemistry taught by Bonnie Hall at Grand View University, Des Moines, USA. This reservation includes Sandbox Reserved 1690 through Sandbox Reserved 1699. |
To get started:
More help: Help:Editing |
Inositol polyphosphate 1-Phosphatase (INPP1) D54A
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ 3.0 3.1 3.2 Dollins DE, Xiong JP, Endo-Streeter S, Anderson DE, Bansal VS, Ponder JW, Ren Y, York JD. A Structural Basis for Lithium and Substrate Binding of an Inositide Phosphatase. J Biol Chem. 2020 Nov 10. pii: RA120.014057. doi: 10.1074/jbc.RA120.014057. PMID:33172890 doi:http://dx.doi.org/10.1074/jbc.RA120.014057