This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Helicase
From Proteopedia
| (2 intermediate revisions not shown.) | |||
| Line 2: | Line 2: | ||
== Function == | == Function == | ||
'''Helicase''' (Hel) is a motor protein which separates nucleic acid strands like DNA double helix or self-annealed RNA. They use ATP hydrolysis for energy. Hel falls into 5 superfamilies (SF1-SF5). Some Hel contain a Helicase and RNase D C terminal | '''Helicase''' (Hel) is a motor protein which separates nucleic acid strands like DNA double helix or self-annealed RNA. They use ATP hydrolysis for energy. Hel falls into 5 superfamilies (SF1-SF5). Some Hel contain a Helicase and RNase D C terminal | ||
| - | Domain (HRDC). The α-thalassemia and mental retardation X-linked syndrome helicase (ATRX ), contains an ATRX-Dnmt3-Dnmt3L (ADD) domain in which many disease-related mutations are found. | + | Domain (HRDC). The α-thalassemia and mental retardation X-linked syndrome helicase (ATRX ), contains an ATRX-Dnmt3-Dnmt3L (ADD) domain in which many disease-related mutations are found.<br /> |
| + | '''ATP-dependent helicase Rho''' is a protein involved in termination of transcription in prokaryotes. Rho binds to the transcription terminator site on single-stranded RNA. Rho forms a ring-shaped hexamer and advances along the mRNA until it reaches the RNA polymerase and causing it to dissociate from the DNA and end transcription.<br /> | ||
| + | '''ATP-dependent helicase RuvB-like 1''' (RuvBL1) or '''TIP49''' is a human protein which forms hexamers. The hexamer forms dodecamer upon association with RuvBL2 or '''TIP48''' and the complex possesses single-stranded DNA-stimulated ATPase and helicase activities. | ||
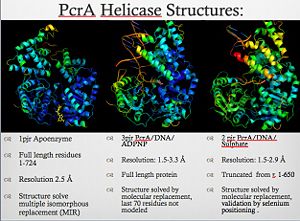
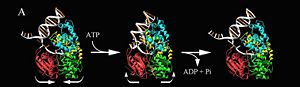
For details of PcrA helicase see<br /> | For details of PcrA helicase see<br /> | ||
| Line 22: | Line 24: | ||
For helicase XPD see<br /> | For helicase XPD see<br /> | ||
*[[XPD Helicase (3CRV)]]<br /> | *[[XPD Helicase (3CRV)]]<br /> | ||
| + | For helicase II or UvrD see<br /> | ||
| + | *[[DNA Repair]]<br /> | ||
| + | For SARS-CoV-2 helicase nsp13 see<br /> | ||
| + | *[[SARS-CoV-2 enzyme Hel]]<br /> | ||
See also<br /> | See also<br /> | ||
*[[Transcription and RNA Processing]] | *[[Transcription and RNA Processing]] | ||
Current revision
| |||||||||||
References
Crystal structure of a DExx box DNA helicase., Subramanya HS, Bird LE, Brannigan JA, Wigley DB, Nature. 1996 Nov 28;384(6607):379-83. PMID:8934527
^ Johnson DS, Bai L, Smith BY, Patel SS, Wang MD (2007). "Single-molecule studies reveal dynamics of DNA unwinding by the ring-shaped t7 helicase". Cell 129 (7): 1299–309. doi:10.1016/j.cell.2007.04.038. PMID 17604719.
^ a b "Researchers solve mystery of how DNA strands separate" (2007-07-03). Retrieved on 2007-07-05.
^ Dumont S, Cheng W, Serebrov V, Beran RK, Tinoco Jr I, Pylr AM, Bustamante C, "RNA Translocation and Unwinding Mechanism of HCV NS3 Helicase and its Coordination by ATP", Nature. 2006 Jan 5; 439: 105-108.
Anand SP, Zheng H, Bianco PR, Leuba SH, Khan SA. DNA helicase activity of PcrA is not required for displacement of RecA protein from DNA or inhibition of RecA-mediated DNA strand exchange. Journal of Bacteriology (2007) 189 (12):4502-4509.
Bird L, Subramanya HS, Wigley DB, "Helicases: a unifying structural theme?", Current Opinion in Structural Biology. 1998 Feb; 8 (1): 14-18.
Betterton MD, Julicher F, "Opening of nucleic-acid double strands by helicases: active versus passive opening.", Physical Review E. 2005 Jan; 71 (1): 011904.
- Sengoku T, Nureki O, Nakamura A, Kobayashi S, Yokoyama S. Structural basis for RNA unwinding by the DEAD-box protein Drosophila Vasa. Cell. 2006 Apr 21;125(2):287-300. PMID:16630817 doi:10.1016/j.cell.2006.01.054
- Sengoku T, Nureki O, Dohmae N, Nakamura A, Yokoyama S. Crystallization and preliminary X-ray analysis of the helicase domains of Vasa complexed with RNA and an ATP analogue. Acta Crystallogr D Biol Crystallogr. 2004 Feb;60(Pt 2):320-2. Epub 2004, Jan 23. PMID:14747711 doi:10.1107/S0907444903025897
Content Donators
Created with the participation of Luis E Ramirez-Tapia, Wayne Decatur.
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Wayne Decatur, Joel L. Sussman