Ribonuclease inhibitor
From Proteopedia
(Difference between revisions)
| (2 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
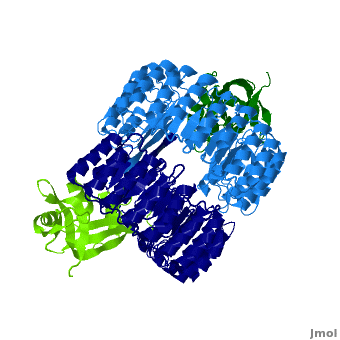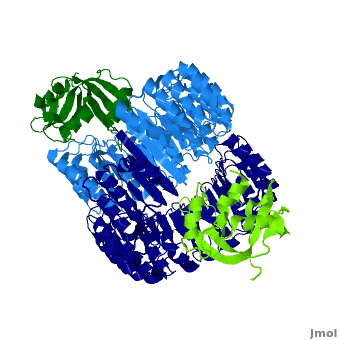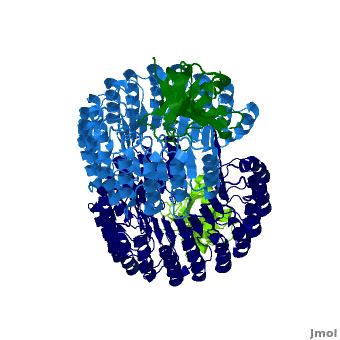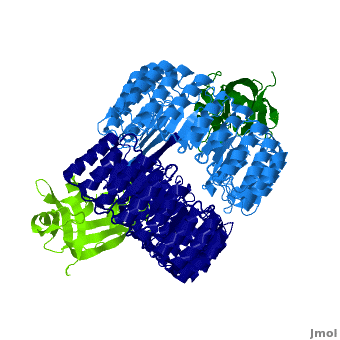
<structuresection load='' size='450' caption='hRI (blue and dark blue) dimer complexed with RNase 1 (light green and dark green), [[1z7x]]' scene='Ribonuclease_inhibitor/1z7x/3' > | <structuresection load='' size='450' caption='hRI (blue and dark blue) dimer complexed with RNase 1 (light green and dark green), [[1z7x]]' scene='Ribonuclease_inhibitor/1z7x/3' > | ||
==Function== | ==Function== | ||
| - | '''Ribonuclease inhibitors (RI)''' are a family of large (~450 residues, ~49 kDa), acidic (pI ~4.7), proteins that bind to and inhibit [http://en.wikipedia.org/wiki/Ribonuclease ribonucleases]. Human RI(hRI) is a major cellular protein, comprising ~0.1% of all cellular protein by weight. <ref>PMID: 11582809</ref> | + | '''Ribonuclease inhibitors (RI)''' or '''Ribonuclease/angiogenin inhibitor''' are a family of large (~450 residues, ~49 kDa), acidic (pI ~4.7), proteins that bind to and inhibit [http://en.wikipedia.org/wiki/Ribonuclease ribonucleases]. Human RI(hRI) is a major cellular protein, comprising ~0.1% of all cellular protein by weight. <ref>PMID: 11582809</ref> |
'''Ribonucleases (RNase)''' are enzymes that degrade RNA and are often [http://en.wikipedia.org/wiki/Cytotoxicity cytotoxic] which gives them chemotherapeutic properties. However, when bound to an RI they are no longer functional. Understanding the mechanism through which RI identifies and binds to RNases will allow scientists to design/modify RNases to evade hRI. In fact, one drug, Onconase (ONC), a ribonuclease from the Northern Leopard Frog ([http://en.wikipedia.org/wiki/Northern_Leopard_Frog Rana pipiens]), is now in Phase III clinical trials as a cancer chemotherapeutic agent <ref>PMID:20173221</ref>. | '''Ribonucleases (RNase)''' are enzymes that degrade RNA and are often [http://en.wikipedia.org/wiki/Cytotoxicity cytotoxic] which gives them chemotherapeutic properties. However, when bound to an RI they are no longer functional. Understanding the mechanism through which RI identifies and binds to RNases will allow scientists to design/modify RNases to evade hRI. In fact, one drug, Onconase (ONC), a ribonuclease from the Northern Leopard Frog ([http://en.wikipedia.org/wiki/Northern_Leopard_Frog Rana pipiens]), is now in Phase III clinical trials as a cancer chemotherapeutic agent <ref>PMID:20173221</ref>. | ||
| Line 49: | Line 49: | ||
As mentioned earlier in the introduction, ribonucleases are cytotoxic. They bind to and chop up RNA. RNase is an endonuclease and is diffusion limited, meaning it acts as fast as susbstrates arrive. RNases exhibit great stability and are often purified by sulfuric acid treatment and then boiling until it is the only surviving macromolecule. RI’s exist to protect cells from rogue RNases. <ref>http://www.pdb.org/pdb/101/motm.do?momID=105</ref> <br> <br> The amphibian RNase ONC is currently in clinical trials, however, human ribonucleases possess advantages over amphibian, namely increased catalytic ability, decreased renal toxicity, and decreased immunogenicity. The RI evasion of an RNase is critical to its chemotherapeutic effectiveness. Genetic engineers have been working on site-specific mutations that either decrease the association constant or increase the disassociation constant. | As mentioned earlier in the introduction, ribonucleases are cytotoxic. They bind to and chop up RNA. RNase is an endonuclease and is diffusion limited, meaning it acts as fast as susbstrates arrive. RNases exhibit great stability and are often purified by sulfuric acid treatment and then boiling until it is the only surviving macromolecule. RI’s exist to protect cells from rogue RNases. <ref>http://www.pdb.org/pdb/101/motm.do?momID=105</ref> <br> <br> The amphibian RNase ONC is currently in clinical trials, however, human ribonucleases possess advantages over amphibian, namely increased catalytic ability, decreased renal toxicity, and decreased immunogenicity. The RI evasion of an RNase is critical to its chemotherapeutic effectiveness. Genetic engineers have been working on site-specific mutations that either decrease the association constant or increase the disassociation constant. | ||
There are many interactions between hRI and RNase 1 providing a multitude of mutagenic options. However, it must be remembered that while changing residues can lead to increased RI evasion, because protein sequence determines structure, and structure determines form changing residues can also lead to decreased catalytic activity. Therefore, altering residues in exchange for RI evasion is a double-edged sword because cytotoxicity must be preserved. It's medically relevant to first understand the interactions between hRI and ribonucleases. Then one can design/modify a ribonuclease to be able to evade hRI while still maintaining functionality—potentially a new way to fight cancer. Researchers at the University of Wisconsin have been working on this very thing. One particular species, “R39D/N67D/N88A/ G89D/R91D RNase 1" has a 5×10^9-fold decrease in affinity for RI, while maintaing nearly wild-type ribonucleolytic activity, conformational stability, and cytotoxicity. <ref>PMID:17350650</ref> The R39D and R91D substitutions sever the hydrogen bonds formed by <scene name='Ribonuclease_inhibitor/Arg91/5'>Arg 91</scene> and <scene name='Ribonuclease_inhibitor/Arg39/1'>Arg 39</scene> that served as electrostatic targeting regions. The aspartates create negative/negative repulsion. This modified RNase provides an interesting example of the future direction and potential of biochemistry and medicine. | There are many interactions between hRI and RNase 1 providing a multitude of mutagenic options. However, it must be remembered that while changing residues can lead to increased RI evasion, because protein sequence determines structure, and structure determines form changing residues can also lead to decreased catalytic activity. Therefore, altering residues in exchange for RI evasion is a double-edged sword because cytotoxicity must be preserved. It's medically relevant to first understand the interactions between hRI and ribonucleases. Then one can design/modify a ribonuclease to be able to evade hRI while still maintaining functionality—potentially a new way to fight cancer. Researchers at the University of Wisconsin have been working on this very thing. One particular species, “R39D/N67D/N88A/ G89D/R91D RNase 1" has a 5×10^9-fold decrease in affinity for RI, while maintaing nearly wild-type ribonucleolytic activity, conformational stability, and cytotoxicity. <ref>PMID:17350650</ref> The R39D and R91D substitutions sever the hydrogen bonds formed by <scene name='Ribonuclease_inhibitor/Arg91/5'>Arg 91</scene> and <scene name='Ribonuclease_inhibitor/Arg39/1'>Arg 39</scene> that served as electrostatic targeting regions. The aspartates create negative/negative repulsion. This modified RNase provides an interesting example of the future direction and potential of biochemistry and medicine. | ||
| - | + | ||
==3D structures of RI== | ==3D structures of RI== | ||
Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| - | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| - | *Ribonuclease inhibitor | ||
| - | + | [[1a4y]] – hRI + RNase 5 – human<br /> | |
| + | [[2bex]] – hRI + nonsecretory RNase<br /> | ||
| + | [[1z7x]], [[2q4g]] - hRI + RNase I<br /> | ||
| + | [[4peq]] - bRI + RNase I - bovine<br /> | ||
| + | [[2bnh]] - pRI - pig<br /> | ||
| + | [[1dfj]] – pRI + bRNase I <br /> | ||
| + | [[3tsr]] – RI + RNAse I – mouse<br /> | ||
| + | [[4per]] - RI + RNase I - chicken<br /> | ||
| - | *Ribonuclease inhibitor·RNase complexes | ||
| - | |||
| - | **[[1dfj]] – pRI + RNase I <br /> | ||
| - | **[[1a4y]] – hRI + RNase 5 – human<br /> | ||
| - | **[[2bex]] – hRI + nonsecretory RNase<br /> | ||
| - | **[[1z7x]], [[2q4g]] - hRI + RNase I<br /> | ||
| - | **[[3tsr]] – RI + RNAse I – mouse<br /> | ||
| - | **[[4peq]] - RI + RNase I - bovine<br /> | ||
| - | **[[4per]] - RI + RNase I - chicken<br /> | ||
| - | |||
| - | }} | ||
==References== | ==References== | ||
<references /> | <references /> | ||
| + | </structuresection> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
Proteopedia Page Contributors and Editors (what is this?)
Abe Weintraub, Michal Harel, Alexander Berchansky, Joel L. Sussman