Sandbox Reserved 963
From Proteopedia
(Difference between revisions)
| (3 intermediate revisions not shown.) | |||
| Line 15: | Line 15: | ||
The structure of WO2 that is described here is that of '''WO2 murine anti-Aβ monoclonal Fab (antigen binding fragment) in the space group P2<sub>1</sub>2<sub>1</sub>2<sub>1</sub> (Form A)'''. There's also another form available for the WO2 antibody, the Form B, but it is not mentioned here because it is crystallized in the space group P2<sub>1</sub>. Thus, in what follows, the different features correspond to the Form A. | The structure of WO2 that is described here is that of '''WO2 murine anti-Aβ monoclonal Fab (antigen binding fragment) in the space group P2<sub>1</sub>2<sub>1</sub>2<sub>1</sub> (Form A)'''. There's also another form available for the WO2 antibody, the Form B, but it is not mentioned here because it is crystallized in the space group P2<sub>1</sub>. Thus, in what follows, the different features correspond to the Form A. | ||
| - | This structure appears to be that of a typical immunoglobulin (Ig) Fab heavy-chain/light-chain heterodimer weighting 52362 Da<ref>http://www.rcsb.org/pdb/explore/explore.do?structureId=3bkm</ref> | + | This structure appears to be that of a typical immunoglobulin (Ig) Fab heavy-chain/light-chain heterodimer weighting 52362 Da<ref>http://www.rcsb.org/pdb/explore/explore.do?structureId=3bkm</ref>. The heavy chain is made up of 252 amino acids while the light chain is made up of 224 amino acids.<ref>http://www.ncbi.nlm.nih.gov/Structure/mmdb/mmdbsrv.cgi?uid=63842</ref> |
| Line 66: | Line 66: | ||
Aβ<sub>1-16</sub> represents the minimal zinc binding domain and contains the entire immunodominant B-cell epitope of Aβ, it is therefore interesting to see how this fragment of Aβ interacts with WO2. | Aβ<sub>1-16</sub> represents the minimal zinc binding domain and contains the entire immunodominant B-cell epitope of Aβ, it is therefore interesting to see how this fragment of Aβ interacts with WO2. | ||
| - | The main residues which closely contact the CDRs of WO2 by sitting within the antigen binding site of WO2 are '''Ala2 to Ser8''' and they stretch 20 Å from the N-terminus to the C-terminus | + | The main residues which closely contact the CDRs of WO2 by sitting within the antigen binding site of WO2 are '''Ala2 to Ser8''' and they stretch 20 Å from the N-terminus to the C-terminus. |
| - | + | The surface area of the Aβ<sub>2-8</sub> structure is 1118 Ų, from which 60% is buried (665 Ų) in the antibody interface. In addition, we note two significant interfaces between Aβ and WO2 : 367 Ų of its surface contacts the heavy chain and 298 Ų contacts the light chain. We notice that residues in the middle of the Aβ<sub>1-16</sub> structure exhibit lower B-factors than atoms at the N- and C- terminus of the Aβ<sub>1-16</sub> peptide, indicating that they are more flexible (since the B-factor, also called the temperature factor, represents the relative vibrational motion of different parts of a structure and atoms with low B-factors belong to a part of the structure quite rigid whereas atoms with high B-factors generally belong to part of a structure that is very flexible[http://en.wikipedia.org/wiki/Debye%E2%80%93Waller_factor]). Phe4 and His6 are completely buried in the Fab interface, each with nearly half of their surface area buried in the V<sub>H</sub> interface and half in the V<sub>L</sub> interface. All other residues are located exclusively at the interface with either the V<sub>H</sub> or the V<sub>L</sub> domains. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | The surface area of the Aβ<sub>2-8</sub> structure is 1118 Ų, from which 60% is buried (665 Ų) in the antibody interface. In addition, we note two significant interfaces between Aβ and WO2 : 367 Ų of its surface contacts the heavy chain and 298 Ų contacts the light chain. We notice | + | |
Residues of the light chain closely contacting Aβ residues include <scene name='60/604482/His_27/2'>His27(D)L</scene>, <scene name='60/604482/Ser27/2'> Ser27(E)L </scene> and <scene name='60/604482/My_first_scene/18'>Tyr32L</scene> from light-chain CDR 1 (L1) and <scene name='60/604482/My_first_scene/7'>Ser92L, Leu93L, Val94L and Leu96L from L3</scene>. | Residues of the light chain closely contacting Aβ residues include <scene name='60/604482/His_27/2'>His27(D)L</scene>, <scene name='60/604482/Ser27/2'> Ser27(E)L </scene> and <scene name='60/604482/My_first_scene/18'>Tyr32L</scene> from light-chain CDR 1 (L1) and <scene name='60/604482/My_first_scene/7'>Ser92L, Leu93L, Val94L and Leu96L from L3</scene>. | ||
| Line 111: | Line 79: | ||
===Details of the close interactions between WO2 and Aβ<sub>2-8</sub>=== | ===Details of the close interactions between WO2 and Aβ<sub>2-8</sub>=== | ||
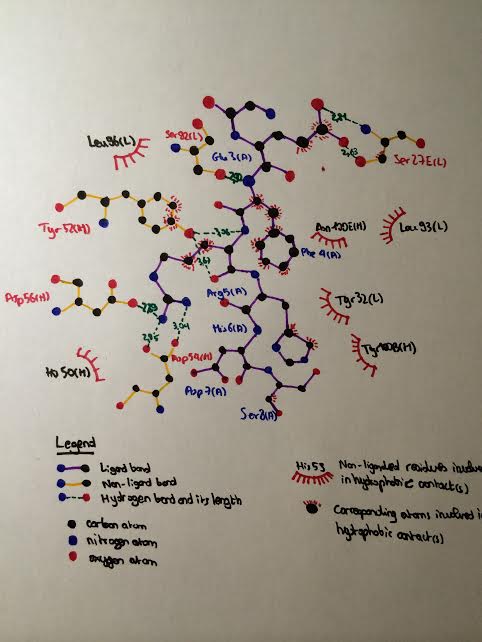
| - | As mentionned previously, the residues of Aβ closely interacting with the CDRs of WO2 extend from Ala2 to Ser8. Let's focus on the interactions of each of these residues with the antibody '''( | + | As mentionned previously, the residues of Aβ closely interacting with the CDRs of WO2 extend from Ala2 to Ser8. Let's focus on the interactions of each of these residues with the antibody '''(see the following LIGPLOT scheme)''' : |
| - | [[Image: | + | [[Image:Ligplot.jpg|right|'''Figure 1''']] |
====Interactions with Ala2==== | ====Interactions with Ala2==== | ||
Ala2 of Aβ<sub>2-8</sub> is recognized by WO2 through a hydrogen bond between its main chain carbonyl group and the amide group of <scene name='60/604482/My_first_scene/4'>Val94L</scene>. | Ala2 of Aβ<sub>2-8</sub> is recognized by WO2 through a hydrogen bond between its main chain carbonyl group and the amide group of <scene name='60/604482/My_first_scene/4'>Val94L</scene>. | ||
| Line 123: | Line 91: | ||
====Interactions with Arg5==== | ====Interactions with Arg5==== | ||
| - | Arg5 is a donor for 5 side chain-side chain hydrogen bonds and 1 main chain-side chain hydrogen bond, involving <scene name='60/604482/Asp54h/1'> Asp54H </scene>, <scene name='60/604482/Asp56h/3'>Asp56H</scene> and <scene name='60/604482/Tyr_25h/1'>Tyr52H</scene> | + | Arg5 is a donor for 5 side chain-side chain hydrogen bonds and 1 main chain-side chain hydrogen bond, involving <scene name='60/604482/Asp54h/1'> Asp54H </scene>, <scene name='60/604482/Asp56h/3'>Asp56H</scene> and <scene name='60/604482/Tyr_25h/1'>Tyr52H</scene> as shown below : |
| - | + | ||
*The side chain of Arg5 forms : | *The side chain of Arg5 forms : | ||
**Two hydrogen bonds with the carboxylate group of <scene name='60/604482/Asp54h/1'> Asp54H </scene> | **Two hydrogen bonds with the carboxylate group of <scene name='60/604482/Asp54h/1'> Asp54H </scene> | ||
| Line 132: | Line 99: | ||
====Interactions with His6==== | ====Interactions with His6==== | ||
| - | His6 contacts both heavy- and light-chain elements with 10 side chain-side chain hydrophobic contacts with Asn100(E)H. Van der Waals contacts have been detected between His6 and <scene name='60/604482/My_first_scene/18'>Tyr32L</scene> of WO2. | + | His6 contacts both heavy- and light-chain elements with 10 side chain-side chain hydrophobic contacts with <scene name='60/604482/Asn100h/1'>Asn100(E)H</scene>. Van der Waals contacts have been detected between His6 and <scene name='60/604482/My_first_scene/18'>Tyr32L</scene> of WO2. |
With Phe4 and Arg5, His6 represents the core of the epitope of Aβ which sits at the heavy-chain/light-chain junction of the CDRs of WO2. | With Phe4 and Arg5, His6 represents the core of the epitope of Aβ which sits at the heavy-chain/light-chain junction of the CDRs of WO2. | ||
| Line 143: | Line 110: | ||
===Comparison of unliganded and liganded WO2 Fab structures=== | ===Comparison of unliganded and liganded WO2 Fab structures=== | ||
| - | Unliganded and liganded structures | + | Unliganded and liganded structures superimpose very closely with an r.m.s.d. (root-mean-square deviation) of 0.3 Å on all Cα atoms (the r.m.s.d. is the measure of the average distance between the atoms of superimposed proteins[http://en.wikipedia.org/wiki/Root-mean-square_deviation]). Even the '''CDRs of liganded and unliganded states are barely distinguishable'''. Except for some small variations (<1 Å) around <scene name='60/604482/Ser27/2'> Ser27(E)L </scene> (L1), <scene name='60/604482/My_first_scene/11'>Lys33H (H1)</scene>, <scene name='60/604482/My_first_scene/16'>Asp54H (H2)</scene> and <scene name='60/604482/My_first_scene/17'>Glu100(C)H (H3)</scene>, there is no substantial change in the CDRs when Aβ binds with WO2. |
Moreover, thanks to temperature-factors analysis, it appears that CDR H1 is less flexible in the liganded structure.<ref>Amyloid-beta-anti-amyloid-beta complex structure reveals an extended conformation in the immunodominant B-cell epitope.,Miles LA, Wun KS, Crespi GA, Fodero-Tavoletti MT, Galatis D, Bagley CJ, Beyreuther K, Masters CL, Cappai R, McKinstry WJ, Barnham KJ, Parker MW J Mol Biol. 2008 Mar 14;377(1):181-92. Epub 2008 Jan 30. PMID:18237744</ref> | Moreover, thanks to temperature-factors analysis, it appears that CDR H1 is less flexible in the liganded structure.<ref>Amyloid-beta-anti-amyloid-beta complex structure reveals an extended conformation in the immunodominant B-cell epitope.,Miles LA, Wun KS, Crespi GA, Fodero-Tavoletti MT, Galatis D, Bagley CJ, Beyreuther K, Masters CL, Cappai R, McKinstry WJ, Barnham KJ, Parker MW J Mol Biol. 2008 Mar 14;377(1):181-92. Epub 2008 Jan 30. PMID:18237744</ref> | ||
| - | + | ||
| Line 154: | Line 121: | ||
The important role of metals in AD is highlighted by a metal chelator, the clioquinol, which reduces amyloid plaques in the brain of AD patients. | The important role of metals in AD is highlighted by a metal chelator, the clioquinol, which reduces amyloid plaques in the brain of AD patients. | ||
| - | The mAb (monoclonal antibody) WO2 recognises the N-terminus of Aβ. This region of Aβ constitutes the immunodominant B-cell epitope of Aβ and lacks T-cell epitopes involved in the toxicity of previous clinical trials. Some experiments showed scientists that the great interest of WO2 lies in the fact that when it binds Aβ, it prevents | + | The mAb (monoclonal antibody) WO2 recognises the N-terminus of Aβ. This region of Aβ constitutes the immunodominant B-cell epitope of Aβ and lacks T-cell epitopes involved in the toxicity of previous clinical trials. Some experiments showed scientists that the great interest of WO2 lies in the fact that when it binds Aβ, it prevents Asp1 and His6 of Aβ to participate in metal coordination, preventing Aβ from aggregating and thus, neutralizing harmful effects of Aβ.<ref>Amyloid-beta-anti-amyloid-beta complex structure reveals an extended conformation in the immunodominant B-cell epitope.,Miles LA, Wun KS, Crespi GA, Fodero-Tavoletti MT, Galatis D, Bagley CJ, Beyreuther K, Masters CL, Cappai R, McKinstry WJ, Barnham KJ, Parker MW J Mol Biol. 2008 Mar 14;377(1):181-92. Epub 2008 Jan 30. PMID:18237744</ref> |
==Related Structures== | ==Related Structures== | ||
Current revision
Anti-amyloid-beta Fab WO2 (Form A, P212121)
| |||||||||||