This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
DAHP synthase
From Proteopedia
(Difference between revisions)
| (13 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
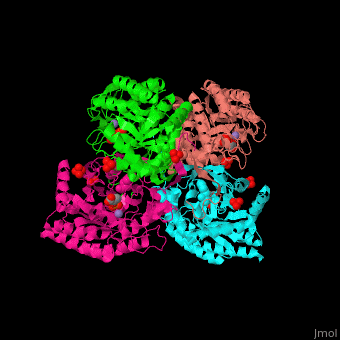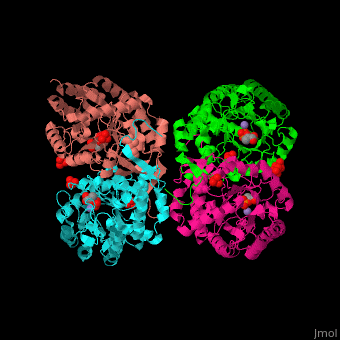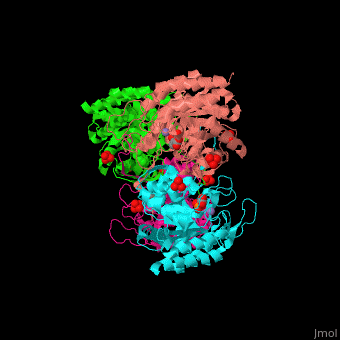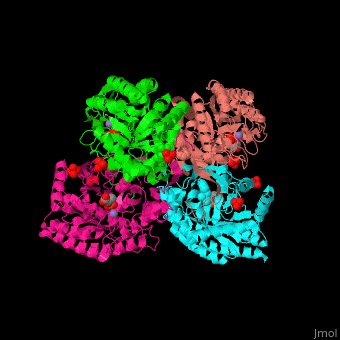
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' caption='E. coli DAHP synthase complex with PEP, phenylalanine, sulfate and Mn+2 ion (PDB code [[1kfl]])' scene='70/708806/Cv/1'> |
== Function == | == Function == | ||
| - | '''DAHP synthase''' or '''3-deoxy-D-arabino-heptulosonate 7-phosphate synthase''' (DAHPS) catalyzes the conversion of phosphoenolpyruvate (PEP) and D-erythrose 4-phosphate to 3-deoxy-D-arabino-heptulosonate 7-phosphate (DAHP) and phosphate. DAHPS is part of the shikimate pathway. DAHPS requires a bivalent metal ion cofactor for normal activity. | + | '''DAHP synthase''' or '''3-deoxy-D-arabino-heptulosonate 7-phosphate synthase''' or '''phospho-2-dehydro-3-deoxyheptonate aldolase''' (DAHPS) catalyzes the conversion of phosphoenolpyruvate (PEP) and D-erythrose 4-phosphate to 3-deoxy-D-arabino-heptulosonate 7-phosphate (DAHP) and phosphate. DAHPS is part of the shikimate pathway. DAHPS requires a bivalent metal ion cofactor for normal activity. <scene name='70/708806/Cv/6'>DAHPS is a tetramer</scene>. DAHPS exhibits feedback inhibition by aromatic amino acids like tyrosine, phenylalanine and tryptophan.<ref>PMID:1682314</ref> |
== Structural highlights == | == Structural highlights == | ||
| - | + | The DAHPS active site is located in a channel at the C-terminal of the enzyme where the <scene name='70/708806/Cv/7'>substrate (PEP)</scene>, <scene name='70/708806/Cv/8'>inhibitor (phenylalanine)</scene> and <scene name='70/708806/Cv/9'>metal ion (Mn+2)</scene> are seen. The bivalent metal is bound to a Cys-X-X-His motif.<ref>PMID:12126632</ref> | |
| - | + | ||
| - | + | ||
== 3D Structures of DAHP synthase == | == 3D Structures of DAHP synthase == | ||
| + | [[DAHP synthase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Stephens CM, Bauerle R. Analysis of the metal requirement of 3-deoxy-D-arabino-heptulosonate-7-phosphate synthase from Escherichia coli. J Biol Chem. 1991 Nov 5;266(31):20810-7. PMID:1682314
- ↑ Shumilin IA, Zhao C, Bauerle R, Kretsinger RH. Allosteric inhibition of 3-deoxy-D-arabino-heptulosonate-7-phosphate synthase alters the coordination of both substrates. J Mol Biol. 2002 Jul 26;320(5):1147-56. PMID:12126632