This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Resolvase
From Proteopedia
(Difference between revisions)
| (10 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
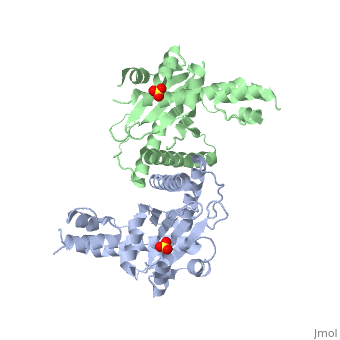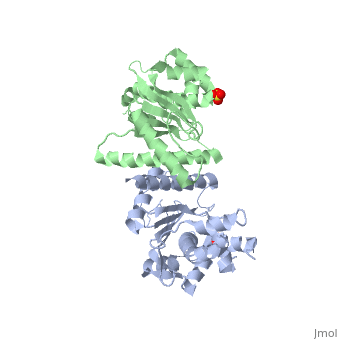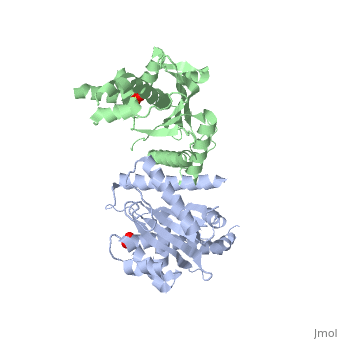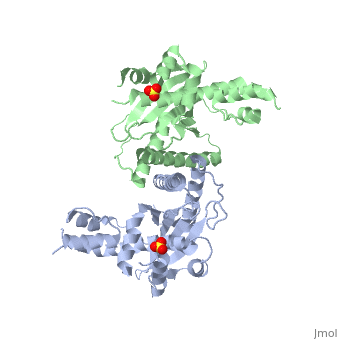
| - | + | <StructureSection load='1kcf' size='340' side='right' caption='Yeast Holliday junction resolvase dimer complex with sulfate, [[1kcf]]' scene='' > | |
== Function == | == Function == | ||
[[Resolvase]] or '''recombinase''' (Rec) is a nuclease which is involved in DNA recombination. According to the binding residue, the recombinases are grouped to Tyr- and Ser-recombinase. | [[Resolvase]] or '''recombinase''' (Rec) is a nuclease which is involved in DNA recombination. According to the binding residue, the recombinases are grouped to Tyr- and Ser-recombinase. | ||
| - | *'''Holliday junction resolvase''' (HJR) resolves 4-way DNA intermediates known as Holliday junctions. Recombination of 2 DNA sites occurs when recombinase binds to the 2 strands. <br /> | + | *'''Holliday junction resolvase''' (HJR) resolves 4-way DNA intermediates known as Holliday junctions. Recombination of 2 DNA sites occurs when recombinase binds to the 2 strands<ref>PMID:19442245</ref>. <br /> |
*'''Tyr-recombinases''' include Cre which recombines loxP sites, FLP and lambda integrase (LamInt). <br /> | *'''Tyr-recombinases''' include Cre which recombines loxP sites, FLP and lambda integrase (LamInt). <br /> | ||
*'''Ser-recombinases''' (S-rec) include gamma-delta resolvase (GDR), Tn3 resolvase and phiC31 integrase.<br /> | *'''Ser-recombinases''' (S-rec) include gamma-delta resolvase (GDR), Tn3 resolvase and phiC31 integrase.<br /> | ||
| - | *'''RadA recombinase''' promotes DNA recombination. Detailed analysis of its <scene name='Resolvase/Cv/2'>dimeric structure</scene> suggests mechanisms for junction isomerization and communication between the two active sites <ref>PMID: | + | *'''RadA recombinase''' promotes DNA recombination. Detailed analysis of its <scene name='Resolvase/Cv/2'>dimeric structure</scene> suggests mechanisms for junction isomerization and communication between the two active sites <ref>PMID:19143611</ref>. <br /> |
| - | *'''Hin-recombinase''' (HRec) is a protein of ''Salmonella''. HRec inverts a 900 base pair DNA segment which contains the promoters of flagellar genes. The inversion changes the these genes' expressions. | + | *'''Hin-recombinase''' (HRec) is a protein of ''Salmonella''. HRec inverts a 900 base pair DNA segment which contains the promoters of flagellar genes. The inversion changes the these genes' expressions<ref>PMID:15454079</ref>. |
==3D structures of resolvase== | ==3D structures of resolvase== | ||
| + | [[Resolvase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | * Holliday junction resolvase | ||
| - | |||
| - | **[[2wiw]], [[2wiz]], [[2wj0]] – AfHJR+DNA – ''Archaeoglobus fulgidus''<br /> | ||
| - | **[[2wcw]], [[2wcz]] – AfHJR (mutant)<br /> | ||
| - | **[[2h8c]] – EcHJR RusA (mutant) + DNA – ''Escherichia coli''<br /> | ||
| - | **[[1gdt]] - EcHJR +DNA<br /> | ||
| - | **[[2h8e]] - EcHJR RusA (mutant)<br /> | ||
| - | **[[1q8r]] – EcHJR RusA<br /> | ||
| - | **[[1hjr]] – EcHJR RuvC<br /> | ||
| - | **[[4ep4]] – TtHJR RuvC – ''Thermus thermophilus''<br /> | ||
| - | **[[4ep5]] – TtHJR RuvC (mutant)<br /> | ||
| - | **[[1zp7]] – HJR RecU – ''Bacillus subtilis''<br /> | ||
| - | **[[1ob8]], [[1ob9]], [[1hh1]] – SsHJR – ''Sulfolobus solfataricus''<br /> | ||
| - | **[[2fco]] – HJR – ''Geobacillus kaustophilus''<br /> | ||
| - | **[[1ipi]], [[1gef]] – HJR – ''Pyrococcus furiosus''<br /> | ||
| - | **[[1kcf]] – HJR – fission yeast<br /> | ||
| - | |||
| - | * Ser-recombinase | ||
| - | |||
| - | **[[2rsl]] - EcGDR<br /> | ||
| - | **[[1hx7]] – EcGDR N-terminal – NMR<br /> | ||
| - | **[[1ght]], [[1res]], [[1ret]], [[1ght]] – EcGDR catalytic domain – NMR<br /> | ||
| - | **[[1gdr]], [[1hx7]] - EcGDR catalytic domain<br /> | ||
| - | **[[2gm4]], [[1zr2]], [[1zr4]], [[1gdt]] – EcGDR+DNA<br /> | ||
| - | **[[2gm5]] – EcGDR (mutant)<br /> | ||
| - | **[[3guv]] – site-specific recombinase – ''Streptococcus pneumoniae''<br /> | ||
| - | **[[2r0q]] – S-rec+DNA – ''Staphylococcus aureus''<br /> | ||
| - | **[[3lhf]] – SsS-rec<br /> | ||
| - | |||
| - | * Tyr-recombinase | ||
| - | |||
| - | **[[3etl]], [[3ew9]], [[3ewa]], [[2i1q]] – MmRadA Rec+AMPPNP+ion – ''Methanococcus maripaludis''<br /> | ||
| - | **[[2b21]] - MmRadA Rec+AMPPNP<br /> | ||
| - | **[[2zub]], [[2zuc]], [[2zud]], [[2dfl]] – SsRadA<br /> | ||
| - | **[[2cvf]], [[2cvh]] – RadB Rec – ''Thermococcus kodakarensis''<br /> | ||
| - | **[[2f1h]], [[2fpl]], [[2fpm]] - MvRec+AMPPNP+K – ''Methanococcus voltae''<br /> | ||
| - | **[[2f1i]] – MvRec+AMPPNP<br /> | ||
| - | **[[2f1j]], [[2fpk]] - MvRec+ADP<br /> | ||
| - | **[[1a0p]] – EcXerd Rec<br /> | ||
| - | |||
| - | * Cre recombinase | ||
| - | |||
| - | **[[3mgv]], [[3c28]], [[3c29]], [[2hof]], [[2hoi]], [[1xns]], [[1xo0]], [[1pvp]], [[1pvq]], [[1pvr]], [[1nzb]], [[1ouq]], [[1q3u]], [[1q3v]], [[1ma7]], [[1kbu]], [[1drg]], [[1f44]], [[2crx]], [[3crx]], [[4crx]], [[5crx]], [[1crx]] – EpCre+DNA – Enterobacteria phage PI<br /> | ||
| - | |||
| - | *Hin recombinase | ||
| - | |||
| - | **[[1hcr]], [[1ijw]], [[1ijj6]], [[1jj8]], [[1jko]], [[1jkp]], [[1jkq]], [[1jkr]] – HRec + DNA – synthetic <br /> | ||
| - | |||
| - | * Integrase | ||
| - | |||
| - | **[[2a3v]] – Rec INTI4+DNA – ''Vibrio cholerae''<br /> | ||
| - | **[[3bvp]] – Integrase N-terminal – Lactococcus phage<br /> | ||
| - | **[[2oxo]] – LamInt binding domain – phage<br /> | ||
| - | **[[1z19]] – EpLamInt core binding +catalytic domain+DNA<br /> | ||
| - | **[[1z1b]], [[1z1g]], [[1p7d]] – EpLamInt+DNA<br /> | ||
| - | **[[1kjk]] – EpLamInt N-terminal – NMR<br /> | ||
| - | **[[1ae9]] – EpLamInt catalytic core<br /> | ||
| - | **[[1p4e]], [[1m6x]] – yFLP (mutant)+DNA – yeast<br /> | ||
| - | **[[1flo]] - yFLP +DNA<br /> | ||
| - | **[[1aih]] – HP1 integrase catalytic domain - bacteriophage<br /> | ||
| - | **[[3nrw]] – INT/site-specific recombinase N-terminal – ''Haloarcula marismortui''<br /> | ||
| - | **[[3jtz]] – INT arm-type binding domain – ''Yersinia pestis''<br /> | ||
| - | **[[3ju0]] – HPI INT arm-type binding domain – ''Pectobacterium atrosepticum''<br /> | ||
| - | **[[2kkp]] – INT SAM-like domain – ''Moorella thermoacetica'' – NMR<br /> | ||
| - | **[[2kkv]] – INT fragment – ''Salmonella enterica'' – NMR<br /> | ||
| - | **[[2khq]] - INT fragment – ''Staphylococcus saprophyticus'' – NMR<br /> | ||
| - | **[[2wcc]] – λINT DBD+ DNA – Enterobacteria phage λ - NMR<br /> | ||
| - | **[[2khv]], [[2kj5]] – INT residues 102-199 – ''Nitrosospira multiformis'' – NMR<br /> | ||
| - | |||
| - | *Recombinase A (RecA) see [[Recombinase A]] | ||
| - | }} | ||
==References== | ==References== | ||
<references /> | <references /> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ West SC. The search for a human Holliday junction resolvase. Biochem Soc Trans. 2009 Jun;37(Pt 3):519-26. doi: 10.1042/BST0370519. PMID:19442245 doi:http://dx.doi.org/10.1042/BST0370519
- ↑ Haldenby S, White MF, Allers T. RecA family proteins in archaea: RadA and its cousins. Biochem Soc Trans. 2009 Feb;37(Pt 1):102-7. doi: 10.1042/BST0370102. PMID:19143611 doi:http://dx.doi.org/10.1042/BST0370102
- ↑ Dhar G, Sanders ER, Johnson RC. Architecture of the hin synaptic complex during recombination: the recombinase subunits translocate with the DNA strands. Cell. 2004 Oct 1;119(1):33-45. PMID:15454079 doi:http://dx.doi.org/10.1016/j.cell.2004.09.010
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Jaime Prilusky