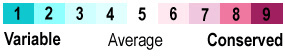This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Introduction to Evolutionary Conservation
From Proteopedia
(Difference between revisions)
| (12 intermediate revisions not shown.) | |||
| Line 104: | Line 104: | ||
<center>{{Template:ColorKey_ConSurf_NoGray}}</center> | <center>{{Template:ColorKey_ConSurf_NoGray}}</center> | ||
| + | ====Sequence Colored by Conservation==== | ||
When ConSurf's colors are applied to the 436 amino acids in the sequence of enolase (based on a multiple sequence alignment containing 150 sequences), this is the result: | When ConSurf's colors are applied to the 436 amino acids in the sequence of enolase (based on a multiple sequence alignment containing 150 sequences), this is the result: | ||
[[Image:4enl consurf150 sequence wb.jpg|400 px|left]] | [[Image:4enl consurf150 sequence wb.jpg|400 px|left]] | ||
{{Clear}} | {{Clear}} | ||
| - | Notice that the conserved residues are scattered around the sequence with no obvious pattern. However, when the same <scene name='Introduction_to_Evolutionary_Conservation/Enolase_with_consurf_colors/1'>colors are applied to the amino acids in the 3D structure</scene>, they form a conserved patch around the catalytic site (marked with a <span style="background:black; color:#00ff00;">''' zinc ion colored green '''</span>. | + | Notice that the conserved residues are scattered around the sequence with no obvious pattern. |
| + | |||
| + | ====3D Structure Colored by Conservation==== | ||
| + | However, when the same <scene name='Introduction_to_Evolutionary_Conservation/Enolase_with_consurf_colors/1'>colors are applied to the amino acids in the 3D structure</scene>, they form a conserved patch around the catalytic site (marked with a <span style="background:black; color:#00ff00;">''' zinc ion colored green '''</span>. | ||
*Show [http://firstglance.jmol.org/fg.htm?mol=http%3A//bioinformatics.org/firstglance/fgij/localPDBFiles/4ENLA_ConSurf_DB_pipe.pdb.gz conservation of enolase in FirstGlance in Jmol] ([[4enl]]). | *Show [http://firstglance.jmol.org/fg.htm?mol=http%3A//bioinformatics.org/firstglance/fgij/localPDBFiles/4ENLA_ConSurf_DB_pipe.pdb.gz conservation of enolase in FirstGlance in Jmol] ([[4enl]]). | ||
| Line 113: | Line 117: | ||
For instructions on how to identify conserved regions of a molecule of interest, and how to show them in Proteopedia (for example with green links), please see [[How to see conserved regions]]. | For instructions on how to identify conserved regions of a molecule of interest, and how to show them in Proteopedia (for example with green links), please see [[How to see conserved regions]]. | ||
| + | |||
| + | ==Expected vs. Unexpected Conservation== | ||
| + | Conservation is '''expected''' for those amino acids that support the 3D structure and functions of a protein. Common examples are listed in the table below. When there is no known structural or functional explanation for conservation of an amino acid, or a cluster of amino acids, the conservation is '''unexpected'''. Unexpected conservation may provide clues for discovering new functions or structural features, e.g. through functional analysis of mutants. | ||
| + | |||
| + | <table class="wikitable"><tr> | ||
| + | <th colspan="2"><center> | ||
| + | Expected Evolutionary Conservation | ||
| + | </center> | ||
| + | </th></tr><tr><th> | ||
| + | Amino Acids | ||
| + | </th><th> | ||
| + | Reason for Conservation | ||
| + | </th></tr><tr><td> | ||
| + | Gly, Pro in turns between helices or beta strands | ||
| + | </td><td> | ||
| + | Required for [[Evolutionary_Conservation#Conservation_for_Domain_Folding|protein domain folding]] | ||
| + | </td></tr><tr><td> | ||
| + | Charged amino acid (Lys, Arg, Asp, Glu) in a salt bridge | ||
| + | </td><td> | ||
| + | Required for [[Salt bridges|protein stability]] | ||
| + | </td></tr><tr><td> | ||
| + | Cys in a disulfide bond | ||
| + | </td><td> | ||
| + | Required for protein stability | ||
| + | </td></tr><tr><td> | ||
| + | N-terminal Met | ||
| + | </td><td> | ||
| + | Start codon for protein synthesis | ||
| + | </td></tr><tr><td> | ||
| + | Amino acids in a large cluster of highly-conserved residues | ||
| + | </td><td> | ||
| + | Required for protein function, e.g. catalytic or binding site | ||
| + | </td></tr></table> | ||
| + | |||
| + | [http://FirstGlance.Jmol.Org FirstGlance in Jmol] makes it easy to locate turns, salt bridges, disulfide bonds, or the N-teminus. In FirstGlance: | ||
| + | * Touch the conserved residue of interest to get its name and sequence number, e.g. Gly236 (in enolase 4enl). | ||
| + | * Use ''Find'' to put yellow halos around the residue of interest, e.g. enter ''Gly236'' in the ''Find'' slot. | ||
| + | ** Turns: Views tab, Secondary Structure. | ||
| + | ** Salt bridges: Tools tab, Salt Bridges. | ||
| + | ** Disulfide bonds: Tools tab, Disulfide Bonds. | ||
| + | ** N terminus: Views tab, N->C Rainbow. You may also wish to check ''Sequence Numbers'' and/or ''Residue Names'' near the bottom of the control panel (upper left panel). | ||
{{Clear}} | {{Clear}} | ||
Current revision
| |||||||||||
See Also
Notes and References
- ↑ MECP2 article in the National Library of Medicine's Genetic Home Reference
- ↑ Advantageous variability will be seen in these cases: 5hmg, 2vaa, 3hi6.