This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox GGC15
From Proteopedia
(Difference between revisions)
| (3 intermediate revisions not shown.) | |||
| Line 5: | Line 5: | ||
== Structure == | == Structure == | ||
| - | '''Human topo 1 is composed of 765 amino acids <ref name="Redinbo" />. The enzyme consist of 4 regions which are the NH2-terminal, core, linker, and COOH-terminal domains<ref name="Redinbo" />. The NH2-terminal is approximately 210 residues long, it is highly charged, disordered, and contains few hydrophobic amino acids<ref name="Redinbo" />. The <scene name='78/781215/04_27_structure_cooh/1'>COOH-terminal</scene> domain is made up of residues 713 to 765 and contains the important amino | + | '''Human topo 1 is composed of 765 amino acids <ref name="Redinbo" />. The enzyme consist of 4 regions which are the NH2-terminal, core, linker, and COOH-terminal domains<ref name="Redinbo" />. The NH2-terminal is approximately 210 residues long, it is highly charged, disordered, and contains few hydrophobic amino acids<ref name="Redinbo" />. The <scene name='78/781215/04_27_structure_cooh/1'>COOH-terminal</scene> domain is made up of residues 713 to 765 and contains the important amino acid <scene name='78/781215/04_27_structure_cooh_with_k/1'>Tyrosine 723</scene><ref name="Redinbo"/>. The location of the active site is at this amino acid<ref name="Redinbo" />. Residues 636 to 712 form the <scene name='78/781215/04_27_structure_linker/1'>Linker domain</scene> and they contribute to the enzyme catalytic activity but are not required<ref name="Redinbo" />. The core is divided into 3 subdomains: <scene name='78/781215/04_27_structure_core_sd1/1'>Core Sudomain 1</scene>[215-232 & 320-433], <scene name='78/781215/04_27_structure_core_sd2/1'>Core Subdomain 2</scene>[233-319], and <scene name='78/781215/04_27_structure_core_sd3/1'>Core Subdomain 3</scene>[434-635]<ref name="Redinbo"/>. The <scene name='78/781215/04_27_structure_core_cooh/1'>core and the COOH-terminal domains</scene> are very important for the catalytic activity<ref name="Redinbo" />.''' |
[[Image:Color Key.jpg]] | [[Image:Color Key.jpg]] | ||
== Active Site & Mechanism Of Action== | == Active Site & Mechanism Of Action== | ||
| Line 13: | Line 13: | ||
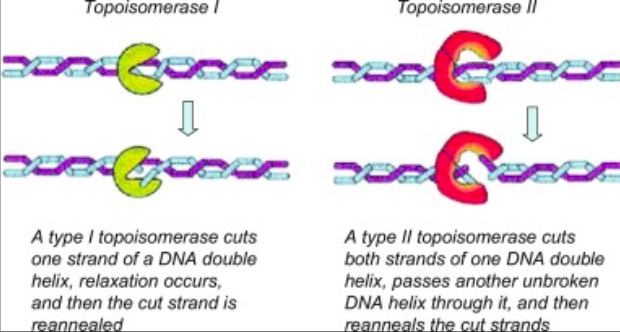
'''The active site of Topo 1 is catalytic and it is the location where the nicking or cutting occurs<ref name="Redinbo" />. The nicking occurs from the trans-esterification of Tyr-723 at a DNA phophodiester bond forming a <scene name='78/781215/04_27_19_active_site_mech/1'>3'-phosphotyrosine covalent enzyme–DNA complex</scene> <ref name="Staker" />. After the DNA is relaxed, the covalent intermediate is reversed when the released 5'-OH of the broken strand reattacks the phosphotyrosine intermediate in a second transesterification reaction<ref name="Staker" />.''' | '''The active site of Topo 1 is catalytic and it is the location where the nicking or cutting occurs<ref name="Redinbo" />. The nicking occurs from the trans-esterification of Tyr-723 at a DNA phophodiester bond forming a <scene name='78/781215/04_27_19_active_site_mech/1'>3'-phosphotyrosine covalent enzyme–DNA complex</scene> <ref name="Staker" />. After the DNA is relaxed, the covalent intermediate is reversed when the released 5'-OH of the broken strand reattacks the phosphotyrosine intermediate in a second transesterification reaction<ref name="Staker" />.''' | ||
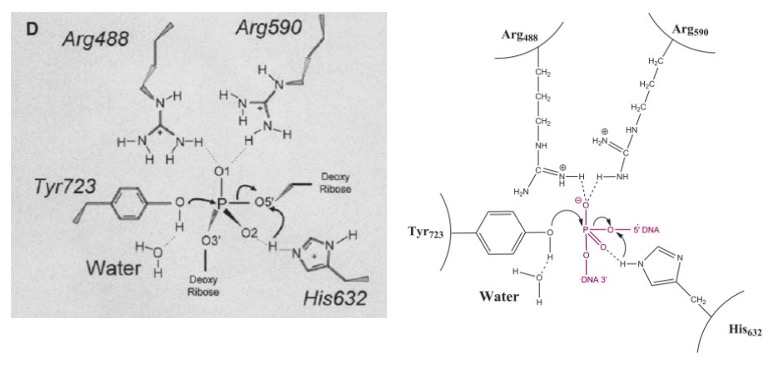
| - | [[Image: | + | [[Image:Active_Site_3_PM.jpg]]<ref name ="Stewart">Stewart, L. (1998). A Model for the Mechanism of Human Topoisomerase I. Science, 279(5356), 1534–1541. https://doi.org/10.1126/science.279.5356.1534</ref> |
[[Image:DNA and Tyrosine.jpg]]<ref name="Stewart" /> | [[Image:DNA and Tyrosine.jpg]]<ref name="Stewart" /> | ||
Current revision
DNA TOPOISOMERASE I
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Staker BL, Hjerrild K, Feese MD, Behnke CA, Burgin AB Jr, Stewart L. The mechanism of topoisomerase I poisoning by a camptothecin analog. Proc Natl Acad Sci U S A. 2002 Nov 26;99(24):15387-92. Epub 2002 Nov 8. PMID:12426403 doi:10.1073/pnas.242259599
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 2.12 2.13 2.14 2.15 2.16 2.17 Redinbo MR, Stewart L, Kuhn P, Champoux JJ, Hol WG. Crystal structures of human topoisomerase I in covalent and noncovalent complexes with DNA. Science. 1998 Mar 6;279(5356):1504-13. PMID:9488644
- ↑ D'yakonov, V. A., Dzhemileva, L. U., & Dzhemilev, U. M. (2017). Advances in the Chemistry of Natural and Semisynthetic Topoisomerase I/II Inhibitors. Studies in Natural Products Chemistry, 21–86. https://doi.org/10.1016/b978-0-444-63929-5.00002-4
- ↑ 4.0 4.1 Stewart, L. (1998). A Model for the Mechanism of Human Topoisomerase I. Science, 279(5356), 1534–1541. https://doi.org/10.1126/science.279.5356.1534
- ↑ 5.0 5.1 Interthal H, Quigley PM, Hol WG, Champoux JJ. The role of lysine 532 in the catalytic mechanism of human topoisomerase I. J Biol Chem. 2004 Jan 23;279(4):2984-92. Epub 2003 Oct 31. PMID:14594810 doi:10.1074/jbc.M309959200