This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Shank protein
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
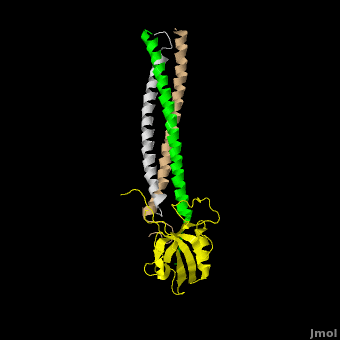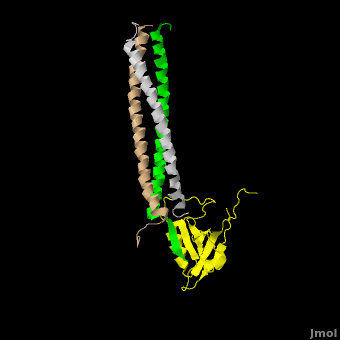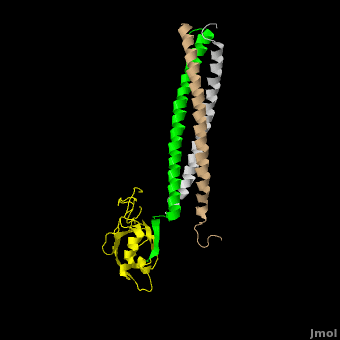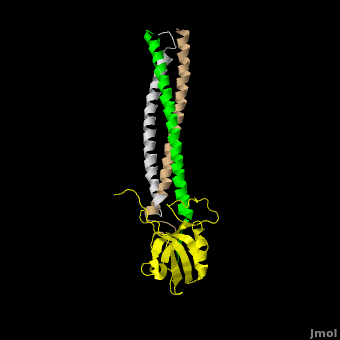
The **canonical PDZ domain** contains 90 amino acids and folds into a compact **globular structure** consisting of a six-stranded β-sandwich flanked by two alpha helices.<ref name="IM"/> βPIX forms a **parallel trimer** via **helical interactions** within its CC domain, and with a **PDZ binding domain** at the C-terminus. Interestingly, only 1 Shank molecule is bound to the CC domain trimer of βPIX in an **asymettric assembly**. (SHOW ZOOMED OUT IN SPACE FILL WITH LONG PART DIRECTLY VERTICAL) The **8-residue PDZ binding domain** (BALL AND STICK AND SPHERE COMBO BURIED MODE) of βPIX forms a number of **hydrogen bonding and hydrophobic interactions** (FIGURE 2A) with the Shank PDZ domain. Shank3-Arg 679 forms the **most critical interaction** with βPIX, tightly binding Glutamate -3. Abolishing this interaction through mutagenesis completely eliminates the assembly. Upon binding of βPIX, the PDZ domain undergoes a significant **conformational change** (OVERVIEW MORPH). Lys 682 undergoes a nearly **11 Angstrom displacement**, ultimately forming a **beta-sheet interaction**, with βPIX residues -4--6, incorporating Shank residues 680 and 681.<ref name="IM"/> | The **canonical PDZ domain** contains 90 amino acids and folds into a compact **globular structure** consisting of a six-stranded β-sandwich flanked by two alpha helices.<ref name="IM"/> βPIX forms a **parallel trimer** via **helical interactions** within its CC domain, and with a **PDZ binding domain** at the C-terminus. Interestingly, only 1 Shank molecule is bound to the CC domain trimer of βPIX in an **asymettric assembly**. (SHOW ZOOMED OUT IN SPACE FILL WITH LONG PART DIRECTLY VERTICAL) The **8-residue PDZ binding domain** (BALL AND STICK AND SPHERE COMBO BURIED MODE) of βPIX forms a number of **hydrogen bonding and hydrophobic interactions** (FIGURE 2A) with the Shank PDZ domain. Shank3-Arg 679 forms the **most critical interaction** with βPIX, tightly binding Glutamate -3. Abolishing this interaction through mutagenesis completely eliminates the assembly. Upon binding of βPIX, the PDZ domain undergoes a significant **conformational change** (OVERVIEW MORPH). Lys 682 undergoes a nearly **11 Angstrom displacement**, ultimately forming a **beta-sheet interaction**, with βPIX residues -4--6, incorporating Shank residues 680 and 681.<ref name="IM"/> | ||
| - | Shank proteins are wedged between scaffolding proteins that are bound to either neurotransmitter receptors or the actin cytoskeleton, making them well positioned to nucleate the underlying structure of the PSD.<ref name="Baron"/> The SAM domain of <scene name='Shank_Family_Proteins/Multimer_opening/1'>Shank3 can oligomerize</scene> (<scene name='Shank_Family_Proteins/Multimer_opening_alt/ | + | Shank proteins are wedged between scaffolding proteins that are bound to either neurotransmitter receptors or the actin cytoskeleton, making them well positioned to nucleate the underlying structure of the PSD.<ref name="Baron"/> The SAM domain of <scene name='Shank_Family_Proteins/Multimer_opening/1'>Shank3 can oligomerize</scene> (<scene name='Shank_Family_Proteins/Multimer_opening_alt/2'>Alternate View</scene>) to form large sheets composed of helical fibers stacked side by side. The proposed sheet structure with radially projecting protein interaction domains, appears to be an ideal architecture for a protein that must contact both membrane and cytoplasmic components at a cell surface.<ref name="Baron"/> Models of this sort validate the importance of Shank3 as master scaffolding proteins and illustrate how slight mutations can disrupt an entire PSD and synaptic function. |
</StructureSection> | </StructureSection> | ||
==Additional Structures of Shank Family Proteins== | ==Additional Structures of Shank Family Proteins== | ||
==References== | ==References== | ||
<references/> | <references/> | ||
Revision as of 02:50, 3 March 2011
| |||||||||||
Additional Structures of Shank Family Proteins
References
- ↑ 1.0 1.1 1.2 Park E, Na M, Choi J, Kim S, Lee JR, Yoon J, Park D, Sheng M, Kim E. The Shank family of postsynaptic density proteins interacts with and promotes synaptic accumulation of the beta PIX guanine nucleotide exchange factor for Rac1 and Cdc42. J Biol Chem. 2003 May 23;278(21):19220-9. Epub 2003 Mar 7. PMID:12626503 doi:10.1074/jbc.M301052200
- ↑ 2.0 2.1 2.2 Baron MK, Boeckers TM, Vaida B, Faham S, Gingery M, Sawaya MR, Salyer D, Gundelfinger ED, Bowie JU. An architectural framework that may lie at the core of the postsynaptic density. Science. 2006 Jan 27;311(5760):531-5. PMID:16439662 doi:311/5760/531
- ↑ 3.0 3.1 3.2 Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, Nygren G, Rastam M, Gillberg IC, Anckarsater H, Sponheim E, Goubran-Botros H, Delorme R, Chabane N, Mouren-Simeoni MC, de Mas P, Bieth E, Roge B, Heron D, Burglen L, Gillberg C, Leboyer M, Bourgeron T. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 2007 Jan;39(1):25-7. Epub 2006 Dec 17. PMID:17173049 doi:ng1933
- ↑ 4.0 4.1 4.2 4.3 Bozdagi O, Sakurai T, Papapetrou D, Wang X, Dickstein DL, Takahashi N, Kajiwara Y, Yang M, Katz AM, Scattoni ML, Harris MJ, Saxena R, Silverman JL, Crawley JN, Zhou Q, Hof PR, Buxbaum JD. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol Autism. 2010 Dec 17;1(1):15. PMID:21167025 doi:10.1186/2040-2392-1-15
- ↑ Garber K. Neuroscience. Autism's cause may reside in abnormalities at the synapse. Science. 2007 Jul 13;317(5835):190-1. PMID:17626859 doi:10.1126/science.317.5835.190
- ↑ Abu-Elneel K, Liu T, Gazzaniga FS, Nishimura Y, Wall DP, Geschwind DH, Lao K, Kosik KS. Heterogeneous dysregulation of microRNAs across the autism spectrum. Neurogenetics. 2008 Jul;9(3):153-61. Epub 2008 Jun 19. PMID:18563458 doi:10.1007/s10048-008-0133-5
- ↑ 7.0 7.1 7.2 7.3 Im YJ, Kang GB, Lee JH, Park KR, Song HE, Kim E, Song WK, Park D, Eom SH. Structural basis for asymmetric association of the betaPIX coiled coil and shank PDZ. J Mol Biol. 2010 Mar 26;397(2):457-66. Epub 2010 Jan 29. PMID:20117114 doi:10.1016/j.jmb.2010.01.048
Proteopedia Page Contributors and Editors (what is this?)
David Canner, Michal Harel, Alexander Berchansky, Joel L. Sussman