This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox 213
From Proteopedia
(→Structure) |
|||
| Line 20: | Line 20: | ||
= Structure = | = Structure = | ||
| + | |||
| + | <StructureSection load='3CLN' size='400' side='left' caption='Structure of calmodulin (PDB entry [[3CLN]])' scene=''> | ||
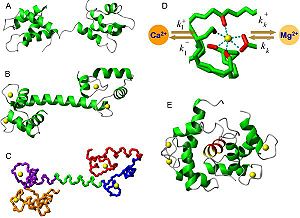
Calmodulin [http://www.rcsb.org/pdb/101/motm_disscussed_entry.do?id=3cln]<ref name="ane">PMID:14527397</ref> is able to bind a large range of target molecules. This property is due to its particularly flexible structure that confers the capacity to change its conformation according to the concentration of calcium in the cell. The calmodulin structure has been determined by NMR. This method reveals that calmodulin is a long molecule which looks like a '''dumbbell''' because it contains '''two globular domains''' (the N-lobe and the C-lobe) linked by a '''flexible α-helix'''. Each lobe contains a pair of helix-loop-helix motifs (called EF-hand) that can bind two Ca<sup>2+</sup> ions. However those lobes do not have the same properties because the C-lobe has higher Ca<sup>2+</sup> affinity than the N-lobe. The two EF-hands are located in the vicinity of each other. Those neighboring sited are very likely to structurally influence each other upon Ca<sup>2+</sup> binding to one of them.<Structure load='1CLL' size='200' frame='true' align='RIGHT' caption='Insert caption here' scene='Insert optional scene name here' /> | Calmodulin [http://www.rcsb.org/pdb/101/motm_disscussed_entry.do?id=3cln]<ref name="ane">PMID:14527397</ref> is able to bind a large range of target molecules. This property is due to its particularly flexible structure that confers the capacity to change its conformation according to the concentration of calcium in the cell. The calmodulin structure has been determined by NMR. This method reveals that calmodulin is a long molecule which looks like a '''dumbbell''' because it contains '''two globular domains''' (the N-lobe and the C-lobe) linked by a '''flexible α-helix'''. Each lobe contains a pair of helix-loop-helix motifs (called EF-hand) that can bind two Ca<sup>2+</sup> ions. However those lobes do not have the same properties because the C-lobe has higher Ca<sup>2+</sup> affinity than the N-lobe. The two EF-hands are located in the vicinity of each other. Those neighboring sited are very likely to structurally influence each other upon Ca<sup>2+</sup> binding to one of them.<Structure load='1CLL' size='200' frame='true' align='RIGHT' caption='Insert caption here' scene='Insert optional scene name here' /> | ||
Revision as of 16:46, 30 December 2011
|
Calmodulin (CaM) for Calcium-Modulated protein is an important protein that intervenes in a wide range of activities inflammation, metabolism, apoptosis, smooth muscle contraction, intracellular movement, short-term and long-term memory, and the immune response. Indeed,it is a small (16.7 kDa = 148 aa) and highly conserved protein that is necessary in all eukaryotic cells because it represents an essential calcium sensor with troponin C its isoform. Calmodulin contains four Ca2+ binding sites and the binding of calcium induces a conformational change in calmodulin that can cause the activation of key enzymes such as kinases or phosphatases proteins (especially phosphorylase kinases) which are not necessarily themselves Ca2+-sensitive and allows a large diversity of cellular response.
Structure
| |||||||||||