This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Chaperonin
From Proteopedia
| Line 34: | Line 34: | ||
== 3D Structures of Chaperonin == | == 3D Structures of Chaperonin == | ||
| - | + | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | |
===Group I=== | ===Group I=== | ||
Revision as of 10:58, 7 March 2013
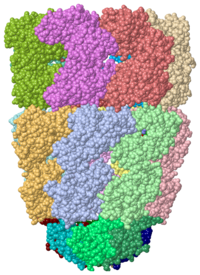
Chaperonins (CPN) are oligomeric proteins that mediate the folding of polypeptide chains. Group I CPN are found in bacteria, chloroplasts and mitochondria. For an introductory overview, see Chaperonins in Wikipedia.
The most characterized CPN are in the GroEL/GroES complex from Escherichia coli and CPN60/CPN10 from Thermus thermophilus. The larger subunit (GroEL, CPN60) contains 3 domains. The apical domain is the one which binds the substrate. Group II CPNs are found in eukaryotic cytosol and archaea. Thermosome is a CPN complex found in archaea. CCT is a CPN complex found in eukarya.
3D Structures of Chaperonin
Updated on 07-March-2013
Group I
Large subunit
3m6c – MtGroEL1 apical domain – Mycobacterium tuberculosis
1sjp, 3rtk – MtGroEL2 residues 42-539
3fbh, 2eu1, 1j4z, 1kpo, 1oel, 1grl, 2yey – EcGroEL (mutant) - Escherichia coli
3e76, 2nwc, 1xck, 1ss8 – EcGroEL
3c9v, 3cau, 2ynj – EcGroEL – Cryo EM
2c7e, 1gr5, 4aaq - EcGroEL (mutant) – Cryo EM
1dk7 - EcGroEL apical domain
1fy9, 1fya, 1kid, 1jon – EcGroEL apical domain (mutant)
1srv – TtCPN60 apical domain - Thermus thermophilus
3osx - CPN60 apical domain – Xenorhabdus nematophila
1iok – CPN60 – Paracoccus denitrificans
Large subunit binary complex
1kp8 - EcGroEL (mutant) + ATP
4aar, 4aas, 4aau, 4ab2, 4ab3 - EcGroEL (mutant) + ATP – Cryo EM
1sx3 - EcGroEL + ATP
1sx4 - EcGroEL + ADP
1mnf – EcGroEL + polypeptide
2cgt – EcGroEL + capsid assbly protein GP31 – EM
1dkd - EcGroEL apical domain + polypeptide
Small subunit
3nx6 – GroES residues 40-134 (mutant) – Xanthomonas oryzae
1wnr – TtCPN10 residues 1-94
1p3h, 1hx5 – MtCPN10
1p82, 1p83 - MtCPN10 residues 1-25 - NMR
1egs - GroES residues 19-27 - NMR
Large + small subunit
1svt, 1pcq - EcGroEL + GroES
1pf9, 1aon - EcGroEL + GroES + ADP
2c7c, 2c7d - EcGroEL + GroES – Cryo EM
1gru - EcGroEL + GroES + ATP + ADP – Cryo EM
1we3 – TtCPN60 + CPN10
1wf4 - TtCPN60 + CPN10 + ADP
Group II
3izh, 3izi, 3izj, 3izk, 3izl, 3izm, 3izn, 3los, 3iyf – MmCPN – Methanococcus maripaludis – EM
3kfb, 3kfe, 3kfk – MmCPN
3j02, 3j03 – MmCPN (mutant) – Cryo EM
3ruq - MmCPN + ADP
3rus - MmCPN (mutant) + ADP
3ruw - MmCPN (mutant) + ADP-AlF3
3ruv - MmCPN (mutant) + ATP analog
1q2v, 1q3r – TkCPN α subunit (mutant) – Thermococcus KS-1
1q3q - TkCPN α subunit (mutant) + AMP-PNP
1q3s - TkCPN α subunit (mutant) + ADP
CCT
2xsm – bCPN CCT – bovine
3p9d, 3p9e, 4d8q, 4d8r – CPN CCT - yeast
3iyg - bCPN CCT – Cryo EM
3ktt - bCPN CCT β subunit – Cryo EM
1gml, 1gn1 - CPN CCT γ subunit apical domain – mouse
Hsp33
3m7m, 1hw7 – EcCPN Hsp33 N-terminal
1xjh - EcCPN Hsp33 C terminal - NMR
1vq0 - CPN Hsp33 – Thermotoga maritima
1vzy - CPN Hsp33 (mutant) – Bacillus subtilis
Thermosome
1a6d - TaTherm α+β subunits – Thermoplasma acidophilum
1a6e - TaTherm α+β subunits + ADP
1ass, 1asx - TaTherm α apical domain
1e0r – TaTherm β apical domain
3ko1 – Therm – Acidianus tengchongensis
3aq1 – Therm – Methanococcoides burtonii
1lep – CPN-10 – Mycobacterium leprae
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Eric Martz, Jaime Prilusky
