This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Isocitrate dehydrogenase
From Proteopedia
| Line 55: | Line 55: | ||
==Inhibition== | ==Inhibition== | ||
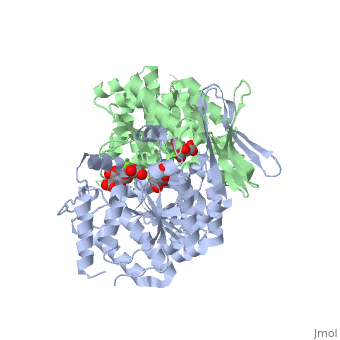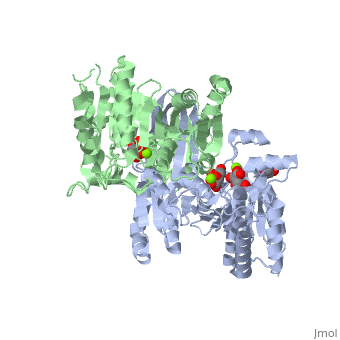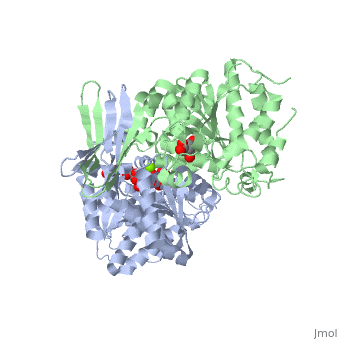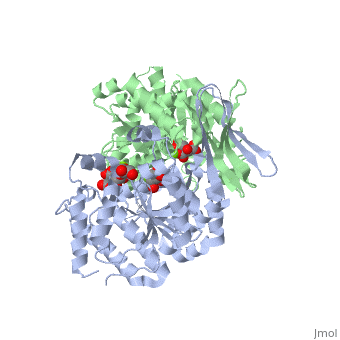
| - | < | + | <scene name='Isocitrate_dehydrogenase/Opening/1'>Structure of Isocitrate Dehydrogenase</scene> ([[1t01]]). |
This step an irreversible reaction. It must be carefully regulated to avoid depletion of isocitrate in the body. The reaction is started by substrate availability which are: isocitrate,Mg2+, NAD+ or NADP+. If these are not present than the process will not carry forward. The reaction is inhibited by the removal of NADH from the presence of the citric acid cycle. The product is also inhibited by ATP feedback. This feedback inhibition is a competitive inhibitor. Since the citric acid cycle is used to produce energy molecules (ATP), an abundance of product will shut down the cycle. | This step an irreversible reaction. It must be carefully regulated to avoid depletion of isocitrate in the body. The reaction is started by substrate availability which are: isocitrate,Mg2+, NAD+ or NADP+. If these are not present than the process will not carry forward. The reaction is inhibited by the removal of NADH from the presence of the citric acid cycle. The product is also inhibited by ATP feedback. This feedback inhibition is a competitive inhibitor. Since the citric acid cycle is used to produce energy molecules (ATP), an abundance of product will shut down the cycle. | ||
Revision as of 13:47, 20 August 2013
| |||||||||||
3D Structures of Isocitrate dehydrogenase
Update February 2013
Isocitrate dehydrogenase
3blx – hIDH - yeast
2uxq – DpIDH – Desulfotalea psychrophila
3dms – IDH – Burkholderia pseudomallei
2e0c, 2dht - StIDH – Sulfolobus tokodaii
2iv0 – IDH – Archaeoglobus fulgidus
1zor – IDH – Thermotoga maritima
2b0t – IDH – Corynebacterium glutamicum
1xgv, 1v94 - ApIDH – Aeropyrum pernix
1pb3, 1sjs, 6icd, 7icd, 3icd – EcIDH – Escherichia coli
1idd, 1idf, 4icd - EcIDH (mutant)
3ah3 – TtIDH – Thermus thermophilus
3us8 – IDH – Sinorhizobium meliloti
4aoy – CtIDH – Clostridium thermocellum
IDH+citrate
2qfw – yIDH+isocitrate
3blv – yIDH+citrate
2uxr – DpIDH+isocitric acid
1itw – AvIDH+isocitrate+Mn – Azotobacter vinelandii
1lwd – IDH+isocitrate+Mn - pig
1pb1, 1p8f – EcIDH+isocitrate
1cw7, 5icd, 8icd - EcIDH +isocitrate+Mg
1cw1 - EcIDH (mutant)+isocitrate+Mn
1gro, 1grp - EcIDH (mutant)+isocitrate+Mg
1hqs – IDH+citrate – Bacillus subtilis
IDH+NADP
1t09 – hIDH+NADP - human
2qfv – yIDH+NADP
2e5m – StIDH+NADP
2cmj – mIDH+NADP – mouse
1tyo - ApIDH+NADP
1j1w – AvIDH+NADP – Azotobacter vinelandii
9icd - EcIDH +NADP
1iso - EcIDH (mutant)+NAD
3mbc - IDH (mutant)+NADP – Corynebacterium glutamicum
IDH+α-ketoglutarate
2qfy - yIDH+α-ketoglutarate
1cw4 – EcIDH (mutant)+α-ketoglutarate
1ika - EcIDH + α-ketoglutarate+Ca
IDH binary complex
3asj – TtIDH + inhbibitor
IDH ternary complexes
3inm – hIDH (mutant)+NADPH+α-ketoglutarate+Ca – human
3map - hIDH+NADP+isocitrate
3mas – hIDH (mutant) +NADP+isocitrate
1t0l – hIDH+NADP+isocitrate+Ca
2qfx - yIDH+NADPH+α-ketoglutarate+Ca
3blw – yIDH+citrate+AMP
2cmv - mIDH+NADP+citrate
2d4v – IDH+NAD+citrate – Acidithiobacillus thiooxidans
2d1c – TtIDH+NADP+citrate
1xkd – ApIDH+NADP+citrate
1hj6 – EcIDH (mutant)+NADP+isopropylmalate+Mg
1idc - EcIDH (mutant)+oxalosuccinate+Mg
1ai3 - EcIDH +isocitrate+Mg+NDP
1ai2, 4aj3, 4aja - EcIDH +isocitrate+NADP+Ca
1ide, 4ajb - EcIDH (mutant)+isocitrate+Mg+NADP
4ajs - EcIDH (mutant)+isocitrate+Mg+ A2P + NMN
4ajc - EcIDH (mutant) +α-ketoglutarate+A2P+Ca
4ajr - EcIDH (mutant) +α-ketoglutarate+ NADP+ Mg
1bl5 - EcIDH +α-ketoglutarate+NADP+Mg
4aou - CtIDH + isocitrate+NADP+Mg
4aov - DpIDH + isocitrate+NADP+Mg
References
- ↑ Xu X, Zhao J, Xu Z, Peng B, Huang Q, Arnold E, Ding J. Structures of human cytosolic NADP-dependent isocitrate dehydrogenase reveal a novel self-regulatory mechanism of activity. J Biol Chem. 2004 Aug 6;279(32):33946-57. Epub 2004 Jun 1. PMID:15173171 doi:10.1074/jbc.M404298200
- ↑ Fedoy AE, Yang N, Martinez A, Leiros HK, Steen IH. Structural and functional properties of isocitrate dehydrogenase from the psychrophilic bacterium Desulfotalea psychrophila reveal a cold-active enzyme with an unusual high thermal stability. J Mol Biol. 2007 Sep 7;372(1):130-49. Epub 2007 Jun 19. PMID:17632124 doi:10.1016/j.jmb.2007.06.040
- ↑ http://en.wikipedia.org/wiki/Isocitrate_dehydrogenase#cite_note-nfr154197.2F32-6
- ↑ Xu X, Zhao J, Xu Z, Peng B, Huang Q, Arnold E, Ding J. Structures of human cytosolic NADP-dependent isocitrate dehydrogenase reveal a novel self-regulatory mechanism of activity. J Biol Chem. 2004 Aug 6;279(32):33946-57. Epub 2004 Jun 1. PMID:15173171 doi:10.1074/jbc.M404298200
- ↑ Xu X, Zhao J, Xu Z, Peng B, Huang Q, Arnold E, Ding J. Structures of human cytosolic NADP-dependent isocitrate dehydrogenase reveal a novel self-regulatory mechanism of activity. J Biol Chem. 2004 Aug 6;279(32):33946-57. Epub 2004 Jun 1. PMID:15173171 doi:10.1074/jbc.M404298200
- ↑ http://en.wikipedia.org/wiki/File:IDHcatalyticmechanism.jpg
- ↑ Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, Li Y, Bhagwat N, Vasanthakumar A, Fernandez HF, Tallman MS, Sun Z, Wolniak K, Peeters JK, Liu W, Choe SE, Fantin VR, Paietta E, Lowenberg B, Licht JD, Godley LA, Delwel R, Valk PJ, Thompson CB, Levine RL, Melnick A. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010 Dec 14;18(6):553-67. Epub 2010 Dec 9. PMID:21130701 doi:10.1016/j.ccr.2010.11.015
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Michael Nobbe, Joel L. Sussman