This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Photosystem II
From Proteopedia
| Line 1: | Line 1: | ||
<StructureSection load='3a0b' size='400' caption='Photosystem II, [[3a0b]]' scene='' > | <StructureSection load='3a0b' size='400' caption='Photosystem II, [[3a0b]]' scene='' > | ||
| - | [[Image:1s5l.gif|250px|left]] | ||
==Background== | ==Background== | ||
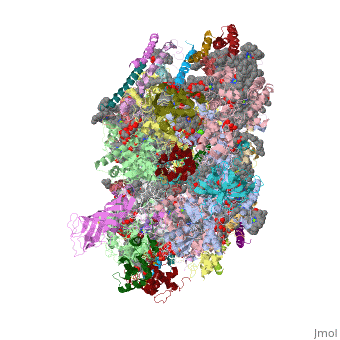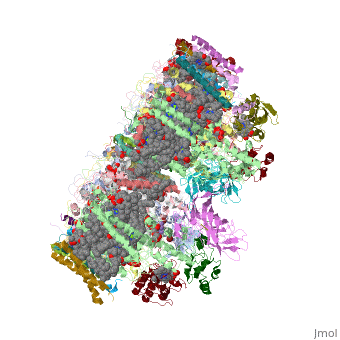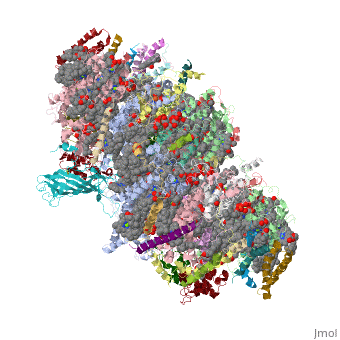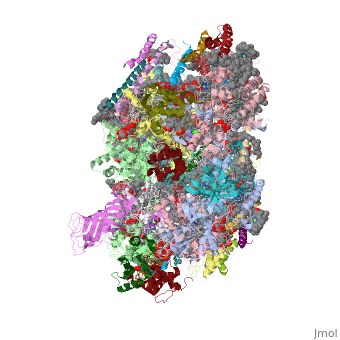
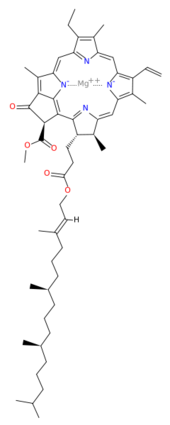
This structure of '''Photosystem II''' was crystallized from the cyanobacteria, ''Thermosynechococcus elongatus'', at 3.0Å <ref>PMID: 16355230</ref> and at 3.50 Å <ref>PMID: 14764885</ref>. PDB codes are [[2axt]] and [[1s5l]], respectively. Cyanobacteria and plants both contain Photosystem II while photosynthetic bacteria contain the bacterial reaction center. This photosynthetic protein complex is associated with a variety of functional ligands. It is a <scene name='Photosystem_II/Psii_dimer/1'>dimer</scene> composed mainly of alpha-helices. Nineteen <scene name='Photosystem_II/Protein_only/1'>subunits</scene> are in each monomer, with multiple extrinsic subunits associated with the oxygen evolving complex missing from this crystallization. Photosystem II is a membrane bound protein complex that in plants is associated with the thylakoid membrane of chloroplasts. <scene name='Photosystem_II/Hydrophobic_polar/1'>Polar and hydrophobic</scene> regions correlate with membrane associated nature of the protein. '''<FONT COLOR="#616D7E">Hydrophobic</FONT>''' helices make up the transmembranal portion, while '''<FONT COLOR="#C031C7">polar</FONT>''' residues are concentrated externally on either side of the membrane. | This structure of '''Photosystem II''' was crystallized from the cyanobacteria, ''Thermosynechococcus elongatus'', at 3.0Å <ref>PMID: 16355230</ref> and at 3.50 Å <ref>PMID: 14764885</ref>. PDB codes are [[2axt]] and [[1s5l]], respectively. Cyanobacteria and plants both contain Photosystem II while photosynthetic bacteria contain the bacterial reaction center. This photosynthetic protein complex is associated with a variety of functional ligands. It is a <scene name='Photosystem_II/Psii_dimer/1'>dimer</scene> composed mainly of alpha-helices. Nineteen <scene name='Photosystem_II/Protein_only/1'>subunits</scene> are in each monomer, with multiple extrinsic subunits associated with the oxygen evolving complex missing from this crystallization. Photosystem II is a membrane bound protein complex that in plants is associated with the thylakoid membrane of chloroplasts. <scene name='Photosystem_II/Hydrophobic_polar/1'>Polar and hydrophobic</scene> regions correlate with membrane associated nature of the protein. '''<FONT COLOR="#616D7E">Hydrophobic</FONT>''' helices make up the transmembranal portion, while '''<FONT COLOR="#C031C7">polar</FONT>''' residues are concentrated externally on either side of the membrane. | ||
Revision as of 06:45, 20 August 2014
| |||||||||||
3D structures of photosystem II
Updated on 20-August-2014
3arc, 3a0b, 3a0h, 4il6 – PSII – Thermosynechococcus vulcanos
3prq, 3prr - TePSII + terbutryn – Thermosynechococcus elongatus
3kzi, 3bz1, 3bz2, 2axt, 1w5c, 1s5l, 1izl, 1ilx, 1fe1, 4fby, 4ixr, 4ixq – TePSII
3zpn - TePSII PSB28 protein
4k7b - PSII extrinsic protein – Chaetoceros gracilis
2y6x – TePSII PSB27 protein
2kvo – SyPSII reaction center PSB28 protein – Synechocystis – NMR
2kmf, 2knd - SyPSII reaction center PSB27 subunit – NMR
2vu4, 1vyk – spPSII PSBP subunit – spinach
1nze - spPSII PSBQ subunit
1v2b - PSII PSBP subunit – tobacco
1fc6, 1fc7, 1fc9, 1fcf – PSII C terminal processing protease – Scenedesmus obliquus
Additional Resources
For additional information, see: Photosynthesis
References
- ↑ Loll B, Kern J, Saenger W, Zouni A, Biesiadka J. Towards complete cofactor arrangement in the 3.0 A resolution structure of photosystem II. Nature. 2005 Dec 15;438(7070):1040-4. PMID:16355230 doi:http://dx.doi.org/10.1038/nature04224
- ↑ Ferreira KN, Iverson TM, Maghlaoui K, Barber J, Iwata S. Architecture of the photosynthetic oxygen-evolving center. Science. 2004 Mar 19;303(5665):1831-8. Epub 2004 Feb 5. PMID:14764885 doi:http://dx.doi.org/10.1126/science.1093087
- ↑ Ferreira KN, Iverson TM, Maghlaoui K, Barber J, Iwata S. Architecture of the photosynthetic oxygen-evolving center. Science. 2004 Mar 19;303(5665):1831-8. Epub 2004 Feb 5. PMID:14764885 doi:http://dx.doi.org/10.1126/science.1093087
Proteopedia Page Contributors and Editors (what is this?)
Emily Forschler, Michal Harel, Ilan Samish, Alexander Berchansky, Eric Martz, Jaime Prilusky, Eran Hodis, Joel L. Sussman, David Canner, Karl Oberholser