This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1085
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
[[Image: Cleavage site of RNase III.png|thumb|right|320px|Fig.3. ''cleavage site of Aa-RNase III'' ]] | [[Image: Cleavage site of RNase III.png|thumb|right|320px|Fig.3. ''cleavage site of Aa-RNase III'' ]] | ||
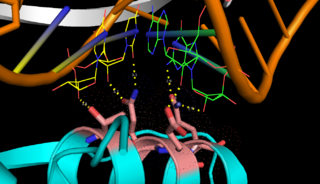
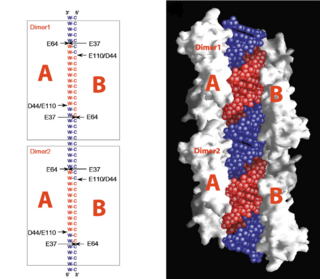
| - | On the basis of the structural and biochemical data, catalytic models were proposed before the structure of a catalytic complex became available. Comparing the structure of Aa-RNase III with the structure of RNA-free Thermotoga maritima RNase III (RNA-free Tm-RNase III, PDB ID code 1O0W)<ref name= Gan>Gan, J., et al., Intermediate states of ribonuclease III in complex with double-stranded RNA. Structure, 2005. 13(10): p. 1435-1442.</ref> shows that there is <scene name='69/699998/Linker_rotation/1'>dramatic rotation</scene> and shift of dsRBD due to RNA binding, and there is a <scene name='69/699998/Linker/2'>flexible linker</scene> KGEMLFD between endoRD and dsRBD leading the rotation happened (''Fig. | + | On the basis of the structural and biochemical data, catalytic models were proposed before the structure of a catalytic complex became available. Comparing the structure of Aa-RNase III with the structure of RNA-free Thermotoga maritima RNase III (RNA-free Tm-RNase III, PDB ID code 1O0W)<ref name= Gan>Gan, J., et al., Intermediate states of ribonuclease III in complex with double-stranded RNA. Structure, 2005. 13(10): p. 1435-1442.</ref> shows that there is <scene name='69/699998/Linker_rotation/1'>dramatic rotation</scene> and shift of dsRBD due to RNA binding, and there is a <scene name='69/699998/Linker/2'>flexible linker</scene> KGEMLFD between endoRD and dsRBD leading the rotation happened (''Fig. 2''). In addition, the two dsRBD domains are spaced apart from each other in a way that allows free rotation of dsRBD around the linker. In vivo data suggested that E110, E37, D44, and E64 are essential for catalysis<ref>Blaszczyk, J., et al., Noncatalytic assembly of ribonuclease III with double-stranded RNA. Structure, 2004. 12(3): p. 457-466.</ref>. This led to the model of the proteins active centers, which can accommodate a dsRNA substrate, each containing two different RNA cleavage sites, <scene name='69/699998/D44_e110_e37_e64/1'>D44/E110 and E37/E64</scene> (pink and red). Specifically, E64 from each partner subunit, along with E37, E110, and D44 are located in the signature motif located at each end of the catalytic valley<ref>Blaszczyk, J., et al., Crystallographic and modeling studies of RNase III suggest a mechanism for double-stranded RNA cleavage. Structure, 2001. 9(12): p. 1225-1236</ref>. ''Fig. 3'' presents a model showing Aa-RNase III cleaving dsRNA. This produces two identical RNA strands, each containing a 2 bp 3' overhang. |
Revision as of 05:32, 24 April 2015
| This Sandbox is Reserved from 15/04/2015, through 15/06/2015 for use in the course "Protein structure, function and folding" taught by Taru Meri at the University of Helsinki. This reservation includes Sandbox Reserved 1081 through Sandbox Reserved 1090. |
To get started:
More help: Help:Editing |
Endoribonuclease III
| |||||||||||
References
- ↑ Lioliou E, Sharma CM, Caldelari I, et al. Global Regulatory Functions of the Staphylococcus aureus Endoribonuclease III in Gene Expression. Hughes D, ed. PLoS Genetics. 2012;8(6):e1002782. doi:10.1371/journal.pgen.1002782
- ↑ Lamontagne, B., et al., The RNase III family: a conserved structure and expanding functions in eukaryotic dsRNA metabolism. Yeast, 2001. 45(191): p. 154-158.
- ↑ Robertson, H.D., Escherichia coli ribonuclease III cleavage sites. Cell, 1982. 30(3): p. 669-672.
- ↑ Grunberg-Manago, M., Messenger RNA stability and its role in control of gene expression in bacteria and phages. Annual review of genetics, 1999. 33(1): p. 193-227.
- ↑ 5.0 5.1 Gan, J., et al., Intermediate states of ribonuclease III in complex with double-stranded RNA. Structure, 2005. 13(10): p. 1435-1442.
- ↑ Blaszczyk, J., et al., Noncatalytic assembly of ribonuclease III with double-stranded RNA. Structure, 2004. 12(3): p. 457-466.
- ↑ Blaszczyk, J., et al., Crystallographic and modeling studies of RNase III suggest a mechanism for double-stranded RNA cleavage. Structure, 2001. 9(12): p. 1225-1236
- ↑ Blaszczyk, J., et al., Noncatalytic assembly of ribonuclease III with double-stranded RNA. Structure, 2004. 12(3): p. 457-466
- ↑ Gan, J., et al., Structural insight into the mechanism of double-stranded RNA processing by ribonuclease III. Cell, 2006. 124(2): p. 355-66