This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Jonathan Lloyd/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 8: | Line 8: | ||
Since much is unknown about ATAD2b we look to its paralogue ATAD2a which has been the focus of many studies. ATAD2a and ATAD2b both share the AAA and bromodomain, in which they are 97% and 74% identical respectively. identical and Based on the observed high sequence similarity with well characterized bromodomains, the ATAD2b bromodomain is expected to recognize acetylated lysine. However since no tests have yet to be done on this additional studies are needed to identify the exact modifications that the ATAD2b bromodomain may recognize. Throughout all bromodomains it seems as though there are very important residues that reside in the binding pocket, Tyr760, Tyr802 and Asn803. These three amino acids are highly conserved and thus must have a major impact in the binding pocket, they are most likely the key to bromodomains binding to acetylated lysine modifications. | Since much is unknown about ATAD2b we look to its paralogue ATAD2a which has been the focus of many studies. ATAD2a and ATAD2b both share the AAA and bromodomain, in which they are 97% and 74% identical respectively. identical and Based on the observed high sequence similarity with well characterized bromodomains, the ATAD2b bromodomain is expected to recognize acetylated lysine. However since no tests have yet to be done on this additional studies are needed to identify the exact modifications that the ATAD2b bromodomain may recognize. Throughout all bromodomains it seems as though there are very important residues that reside in the binding pocket, Tyr760, Tyr802 and Asn803. These three amino acids are highly conserved and thus must have a major impact in the binding pocket, they are most likely the key to bromodomains binding to acetylated lysine modifications. | ||
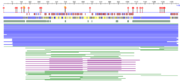
| - | [[Image:table 2.png |thumb| | + | [[Image:table 2.png |thumb| The entire ATAD2b protein sequenced using prediction software. Diamonds represent protein binding, circles represent nucleosome binding. Blue boxes are alpha helix, red boxes beta sheet.]] |
== Sequence Analysis == | == Sequence Analysis == | ||
A BLAST search of the bromodomain in ATAD2b yields many other bromodomain-containing proteins. In the table below are 8 other proteins that contain a bromodomain. Their sequences were aligned to ATAD2b using Clustal Omega and show the percent identity and similarity to ATAD2b. You can see the conserved residues throughout all the proteins (marked with *), there are 13 out of the 71 residues or 18.3%. Many of the identical residues are a result of these 13 residues showing how highly conserved they are. They have very important function in the bromodomain which is why it would be expected to see the same residues in all bromodomains. The closest protein to ATAD2b is ATAD2a which is no surprise considering they are paralogues of each other. | A BLAST search of the bromodomain in ATAD2b yields many other bromodomain-containing proteins. In the table below are 8 other proteins that contain a bromodomain. Their sequences were aligned to ATAD2b using Clustal Omega and show the percent identity and similarity to ATAD2b. You can see the conserved residues throughout all the proteins (marked with *), there are 13 out of the 71 residues or 18.3%. Many of the identical residues are a result of these 13 residues showing how highly conserved they are. They have very important function in the bromodomain which is why it would be expected to see the same residues in all bromodomains. The closest protein to ATAD2b is ATAD2a which is no surprise considering they are paralogues of each other. | ||
Revision as of 22:50, 9 December 2015
ATAD2b
| |||||||||||