This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Carlos Henrique Passos/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 39: | Line 39: | ||
There are show two <scene name='78/786632/Disulfide_bridges/1'>disulfide bridges</scene> on this protein. These bridges are made by the presence of cysteines on the central beta-sheet, on the positions 34 and 178 (Cys34 and Cys178). One disulfide bridge is the <scene name='78/786632/Cys34_cys123_bridge/1'>Cys34-Cys123</scene> interactions, and the other is <scene name='78/786632/Cys178_cys249_bridge/1'>Cys178-Cys249</scene> <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. The fifth <scene name='78/786632/Free_cys_73/1'>Cysteine 73</scene> is not participating in any disulfide bridge. | There are show two <scene name='78/786632/Disulfide_bridges/1'>disulfide bridges</scene> on this protein. These bridges are made by the presence of cysteines on the central beta-sheet, on the positions 34 and 178 (Cys34 and Cys178). One disulfide bridge is the <scene name='78/786632/Cys34_cys123_bridge/1'>Cys34-Cys123</scene> interactions, and the other is <scene name='78/786632/Cys178_cys249_bridge/1'>Cys178-Cys249</scene> <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. The fifth <scene name='78/786632/Free_cys_73/1'>Cysteine 73</scene> is not participating in any disulfide bridge. | ||
| - | ---- | ||
| - | ---- | ||
'''Hidrogen Bonds''' | '''Hidrogen Bonds''' | ||
| Line 47: | Line 45: | ||
The folding of the protein is stabilized by a total of 130 hydrogen bonds, of which 55 are made between the main chain and side chain functional groups, and 75 are made between side chains. If we include the water molecules, the hydrogen bonds interactions rise up to 191 interactions. <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. | The folding of the protein is stabilized by a total of 130 hydrogen bonds, of which 55 are made between the main chain and side chain functional groups, and 75 are made between side chains. If we include the water molecules, the hydrogen bonds interactions rise up to 191 interactions. <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. | ||
| + | |||
| + | ---- | ||
| + | ---- | ||
===Active Site and Substrate Recognition=== | ===Active Site and Substrate Recognition=== | ||
| Line 62: | Line 63: | ||
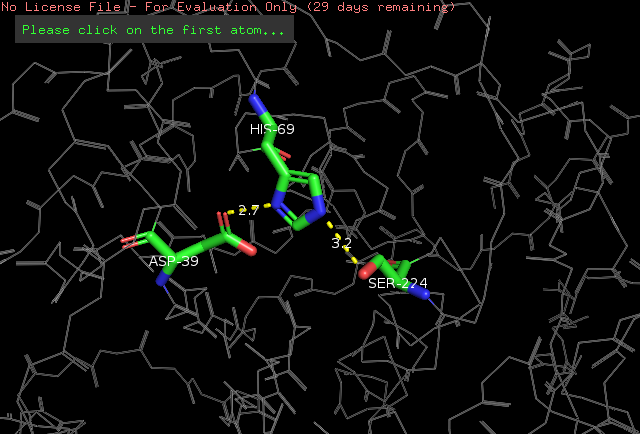
The active site of the protein is located on the central beta-sheet (as described above). The Asp39 aminoacid accepts two hydrogen bonds, on the carbonyl oxygen, from Val95 and Leu96, in a way that stabilizes this important residue <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. | The active site of the protein is located on the central beta-sheet (as described above). The Asp39 aminoacid accepts two hydrogen bonds, on the carbonyl oxygen, from Val95 and Leu96, in a way that stabilizes this important residue <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>. | ||
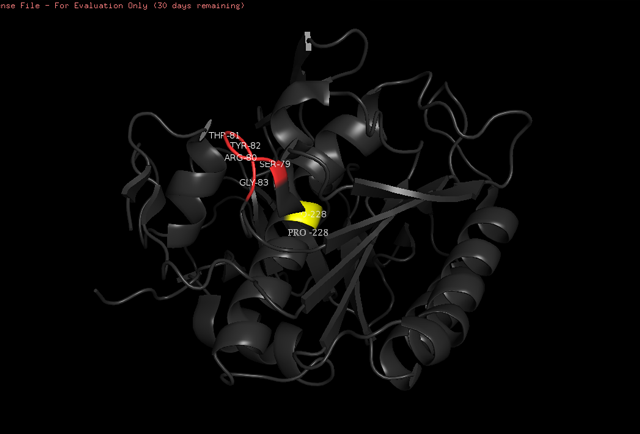
The <scene name='78/786632/Substrate_recognition/1'>susbtrate recognition</scene> are made by two peptide chains, one that englobes aminoacids ranging from 99 to 104 positions, and the other made of aminoacids ranging from 132 to 136 positions, which can be viewed <scene name='78/786632/Substrate_recognition/2'>here</scene>. The colors indicate where there is a region with alpha-helix (in pink), beta-sheet (in gold), or loop (in white). | The <scene name='78/786632/Substrate_recognition/1'>susbtrate recognition</scene> are made by two peptide chains, one that englobes aminoacids ranging from 99 to 104 positions, and the other made of aminoacids ranging from 132 to 136 positions, which can be viewed <scene name='78/786632/Substrate_recognition/2'>here</scene>. The colors indicate where there is a region with alpha-helix (in pink), beta-sheet (in gold), or loop (in white). | ||
| + | |||
| + | ---- | ||
| + | ---- | ||
===Ligands=== | ===Ligands=== | ||
| Line 72: | Line 76: | ||
he other calcium binding site is less well-defined <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>, and are formed by the amino acids Thr16 and Asp260, as shown <scene name='78/786632/Ca_binding_site_2/1'>here</scene>. Calcium is not represented in this scene. | he other calcium binding site is less well-defined <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref>, and are formed by the amino acids Thr16 and Asp260, as shown <scene name='78/786632/Ca_binding_site_2/1'>here</scene>. Calcium is not represented in this scene. | ||
| - | + | ---- | |
| + | ---- | ||
Revision as of 00:34, 18 June 2018
==Proteinase K==
| |||||||||||
References
- ↑ PDB 2ID8: Wang, Jiawei, Miroslawa Dauter, and Zbigniew Dauter. "What can be done with a good crystal and an accurate beamline?." Acta Crystallographica Section D: Biological Crystallography 62.12 (2006): 1475-1483. Available: https://www.ncbi.nlm.nih.gov/pubmed/17139083
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Cronier, Sabrina, et al. "Detection and characterization of proteinase K-sensitive disease-related prion protein with thermolysin." Biochemical Journal 416.2 (2008): 297-305. Available online: http://www.biochemj.org/content/416/2/297