This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Carlos Henrique Passos/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 51: | Line 51: | ||
'''Active Site and Substrate Recognition''' | '''Active Site and Substrate Recognition''' | ||
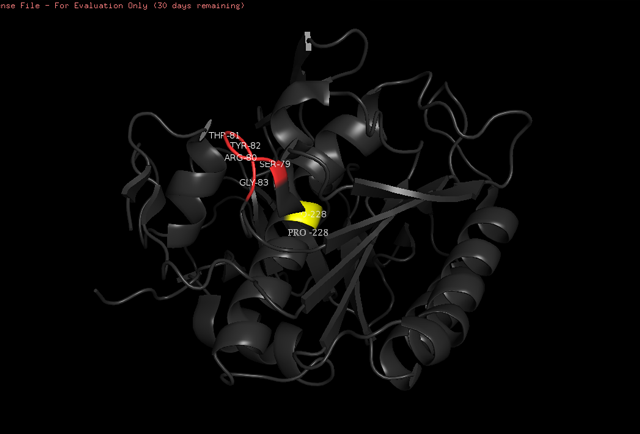
| - | The <scene name='78/786632/Active_site/3'>catalytic site</scene> of Proteinase K is made by a | + | The <scene name='78/786632/Active_site/3'>catalytic site</scene> of Proteinase K is made by a triad of amino acids, <scene name='78/786632/Asp39/2'>Asp39</scene>, <scene name='78/786632/His69/2'>His69</scene> and <scene name='78/786632/Ser224/2'>Ser224</scene> <ref>Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x</ref> (hence it is a serine protease). The pink color indicates that these amino acids are all polars. See more about Proteinase K function in the Function section below. |
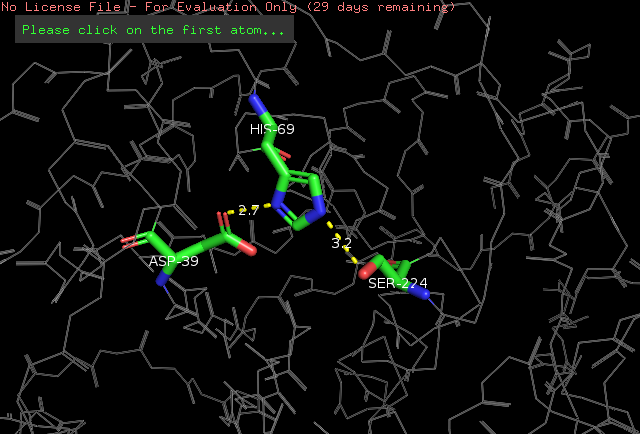
The amino acids side chains of the catalytic triad are stabilized by hydrogen bonds, as demonstrated by a simulation done in the PyMOL software. We know that these are hydrogen bonds because the distance between the atoms participating in the ligation is around 2.8 angstroms. See the simulation: | The amino acids side chains of the catalytic triad are stabilized by hydrogen bonds, as demonstrated by a simulation done in the PyMOL software. We know that these are hydrogen bonds because the distance between the atoms participating in the ligation is around 2.8 angstroms. See the simulation: | ||
Revision as of 01:42, 18 June 2018
==Proteinase K==
| |||||||||||
References
- ↑ PDB 2ID8: Wang, Jiawei, Miroslawa Dauter, and Zbigniew Dauter. "What can be done with a good crystal and an accurate beamline?." Acta Crystallographica Section D: Biological Crystallography 62.12 (2006): 1475-1483. Available: https://www.ncbi.nlm.nih.gov/pubmed/17139083
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Betzel, Christian, Gour P. PAL, and Wolfram SAENGER. "Three‐dimensional structure of proteinase K at 0.15‐nm resolution." The FEBS Journal 178.1 (1988): 155-171. Available: https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1432-1033.1988.tb14440.x
- ↑ Cronier, Sabrina, et al. "Detection and characterization of proteinase K-sensitive disease-related prion protein with thermolysin." Biochemical Journal 416.2 (2008): 297-305. Available online: http://www.biochemj.org/content/416/2/297