User:Ashley Crotteau/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 12: | Line 12: | ||
===SET Domain=== | ===SET Domain=== | ||
| - | The human lysine methyltransferase (HKMT) SET7/9 is 366 amino acids long. The overall structure looks like a dimer although it acts as a monomer. The structure is composed of the ΔSET7/9 domain. The ΔSET7/9 consists of the SET domain along with the pre- and post-SET regions.<ref name="Schubert" /><ref name="Yeates" /> The pre- and post-SET regions are adjacent to SET domain and are cysteine rich.<ref name="Schubert" /><ref name="Yeates" /> The pre-SET cysteine region is located near the N-terminal where the post-SET region is located near the C-terminal of the domain.<ref name="Schubert" /><ref name="Yeates" /> These regions are said to play an important role in substrate recognition and enzymatic activity.<ref name="Schubert" /><ref name="Yeates" /> | + | The <scene name='81/811707/Overall_structure/1'>human lysine methyltransferase </scene>(HKMT) SET7/9 is 366 amino acids long. The overall structure looks like a dimer although it acts as a monomer. The structure is composed of the ΔSET7/9 domain. The ΔSET7/9 consists of the SET domain along with the pre- and post-SET regions.<ref name="Schubert" /><ref name="Yeates" /> The pre- and post-SET regions are adjacent to SET domain and are cysteine rich.<ref name="Schubert" /><ref name="Yeates" /> The pre-SET cysteine region is located near the N-terminal where the post-SET region is located near the C-terminal of the domain.<ref name="Schubert" /><ref name="Yeates" /> These regions are said to play an important role in substrate recognition and enzymatic activity.<ref name="Schubert" /><ref name="Yeates" /> |
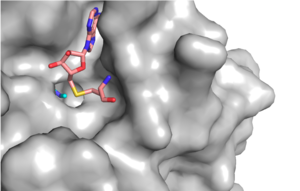
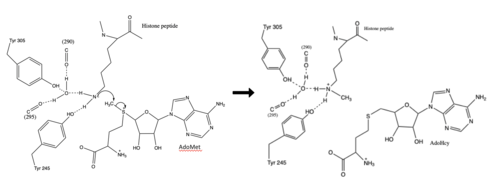
The SET domain is mostly defined by <scene name='81/811707/Variable_knot/2'>turns and loops</scene> with the few <scene name='81/811707/Beta_sheets/3'>antiparallel β-sheets</scene>.<ref name="Schubert" /> <scene name='81/811707/Beta-hairpin/2'>Residues 337-349</scene> form a β-hairpin that sticks out at a right angle to the surface of the enzyme.<ref name="Xiao" /> The following three residues (<scene name='81/811707/Sharp_bend/2'>350-352</scene>) accommodate a sharp bend in the peptide chain and the end of the protein adopts an <scene name='81/811707/C-term_alpha_helix/2'>α-helical conformation</scene>.<ref name="Xiao" /> The two most defining features of the SET domain are the C-terminal tyrosine and the knot-like fold. These two components have been recognized to be essential for <scene name='81/811708/Sam_structure/3'>S-adenosyl-L-methionine</scene> (SAM) binding and catalysis.<ref name="Schubert" /> <ref name="Yeates" /> <ref name="Huang" /> The knot-like fold contains the binding sites for the cofactor SAM and the peptide substrate.<ref name="Licciardello" /> | The SET domain is mostly defined by <scene name='81/811707/Variable_knot/2'>turns and loops</scene> with the few <scene name='81/811707/Beta_sheets/3'>antiparallel β-sheets</scene>.<ref name="Schubert" /> <scene name='81/811707/Beta-hairpin/2'>Residues 337-349</scene> form a β-hairpin that sticks out at a right angle to the surface of the enzyme.<ref name="Xiao" /> The following three residues (<scene name='81/811707/Sharp_bend/2'>350-352</scene>) accommodate a sharp bend in the peptide chain and the end of the protein adopts an <scene name='81/811707/C-term_alpha_helix/2'>α-helical conformation</scene>.<ref name="Xiao" /> The two most defining features of the SET domain are the C-terminal tyrosine and the knot-like fold. These two components have been recognized to be essential for <scene name='81/811708/Sam_structure/3'>S-adenosyl-L-methionine</scene> (SAM) binding and catalysis.<ref name="Schubert" /> <ref name="Yeates" /> <ref name="Huang" /> The knot-like fold contains the binding sites for the cofactor SAM and the peptide substrate.<ref name="Licciardello" /> | ||
Revision as of 17:47, 5 April 2019
H. sapiens Lysine Methyltransferase, SET 7/9
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Schubert HL, Blumenthal RM, Cheng X. Many paths to methyltransfer: a chronicle of convergence. Trends Biochem Sci. 2003 Jun;28(6):329-35. PMID:12826405
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Yeates TO. Structures of SET domain proteins: protein lysine methyltransferases make their mark. Cell. 2002 Oct 4;111(1):5-7. PMID:12372294
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378
- ↑ 4.0 4.1 Huang S, Shao G, Liu L. The PR domain of the Rb-binding zinc finger protein RIZ1 is a protein binding interface and is related to the SET domain functioning in chromatin-mediated gene expression. J Biol Chem. 1998 Jun 26;273(26):15933-9. PMID:9632640
- ↑ 5.0 5.1 doi: https://dx.doi.org/10.1016/C2014-0-02189-2
- ↑ https://en.wikipedia.org/wiki/SET_domain#Structure
Student Contributors
Ashley Crotteau
Parker Hiday
Lauren Allman