This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1700
From Proteopedia
(Difference between revisions)
| Line 46: | Line 46: | ||
</jmol> | </jmol> | ||
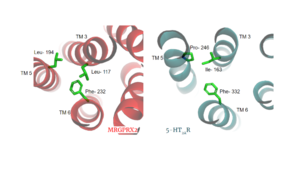
| - | LLF | + | Another motif found in most, but not all, A family GPCR’s is the PIF motif. The three residues are found on transmembrane helices 5, 3, and 6, respectively. In MRGPRX2, the PIF motif is changed to LLF residues. Figure 3 shows the conserved PIF motif on 5HT2AR, compared to the LLF motif on MRGPRX2, found at <scene name='90/904305/Llf_motif/3'>Leu-117, Leu-194, and Phe-232</scene> on transmembrane helices 5, 3, and 6, respectively. This contributes to shifting helix 6 towards helix 3, and contributes to the tighter packing of helices and therefore a more surface-level ligand binding site. |
| + | |||
| + | [[Image:LLF.png|300px|right|thumb|Figure 3. Conserved PIF motif in 5HT2AR (teal) compared to the LLF motif found in MRGPRX2 (red). Transmembrane helices and residues are numbered and labeled to show how this structural change shifts the orientation of the helices.]] | ||
| + | |||
==== DRY/ ERC Motif ==== | ==== DRY/ ERC Motif ==== | ||
Revision as of 00:32, 29 March 2022
| This Sandbox is Reserved from February 28 through September 1, 2022 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1700 through Sandbox Reserved 1729. |
To get started:
More help: Help:Editing |
MRGPRX2 Human Itch G-Protein Coupled Receptor (GPCR)
| |||||||||||
References
- ↑ Hauser AS, Attwood MM, Rask-Andersen M, Schioth HB, Gloriam DE. Trends in GPCR drug discovery: new agents, targets and indications. Nat Rev Drug Discov. 2017 Dec;16(12):829-842. doi: 10.1038/nrd.2017.178. Epub, 2017 Oct 27. PMID:29075003 doi:http://dx.doi.org/10.1038/nrd.2017.178
- ↑ Basith S, Cui M, Macalino SJY, Park J, Clavio NAB, Kang S, Choi S. Exploring G Protein-Coupled Receptors (GPCRs) Ligand Space via Cheminformatics Approaches: Impact on Rational Drug Design. Front Pharmacol. 2018 Mar 9;9:128. doi: 10.3389/fphar.2018.00128. eCollection, 2018. PMID:29593527 doi:http://dx.doi.org/10.3389/fphar.2018.00128
- ↑ 3.0 3.1 3.2 Cao C, Kang HJ, Singh I, Chen H, Zhang C, Ye W, Hayes BW, Liu J, Gumpper RH, Bender BJ, Slocum ST, Krumm BE, Lansu K, McCorvy JD, Kroeze WK, English JG, DiBerto JF, Olsen RHJ, Huang XP, Zhang S, Liu Y, Kim K, Karpiak J, Jan LY, Abraham SN, Jin J, Shoichet BK, Fay JF, Roth BL. Structure, function and pharmacology of human itch GPCRs. Nature. 2021 Dec;600(7887):170-175. doi: 10.1038/s41586-021-04126-6. Epub 2021, Nov 17. PMID:34789874 doi:http://dx.doi.org/10.1038/s41586-021-04126-6
- ↑ 4.0 4.1 4.2 Yang F, Guo L, Li Y, Wang G, Wang J, Zhang C, Fang GX, Chen X, Liu L, Yan X, Liu Q, Qu C, Xu Y, Xiao P, Zhu Z, Li Z, Zhou J, Yu X, Gao N, Sun JP. Structure, function and pharmacology of human itch receptor complexes. Nature. 2021 Dec;600(7887):164-169. doi: 10.1038/s41586-021-04077-y. Epub 2021, Nov 17. PMID:34789875 doi:http://dx.doi.org/10.1038/s41586-021-04077-y
- ↑ Trzaskowski B, Latek D, Yuan S, Ghoshdastider U, Debinski A, Filipek S. Action of molecular switches in GPCRs--theoretical and experimental studies. Curr Med Chem. 2012;19(8):1090-109. doi: 10.2174/092986712799320556. PMID:22300046 doi:http://dx.doi.org/10.2174/092986712799320556