This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
BASIL2022GV3R8E
From Proteopedia
| Line 26: | Line 26: | ||
[[Image:FinalGel_2.png|250px|]] | [[Image:FinalGel_2.png|250px|]] | ||
== Project Implications == | == Project Implications == | ||
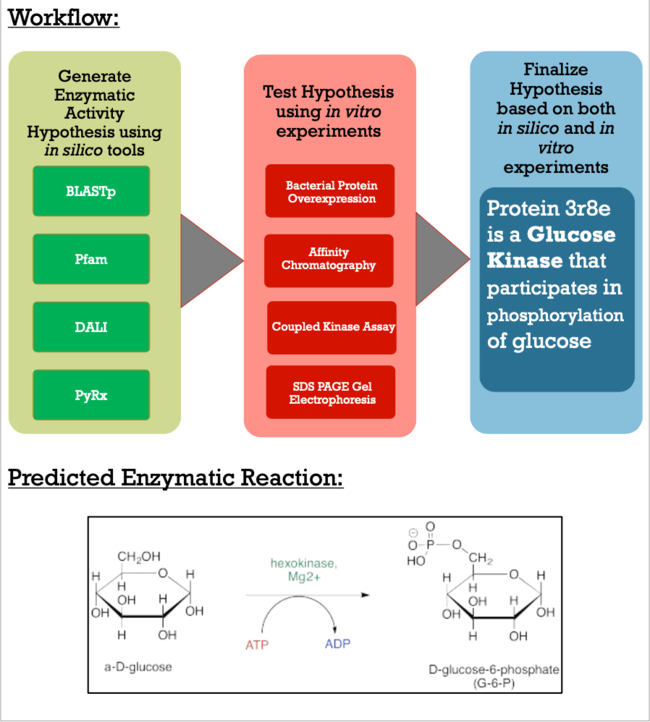
| - | The goal of this project is to explore the | + | The goal of this project is to explore the techniques it takes to characterize a putative kinase. To do this, we became familiar with online alignment, structure, and function tools, paired with a variety of in vitro lab experiments, including bacterial protein overexpression, affinity chromatography, coupled kinase assays, and SDS PAGE. These techniques can be used to help characterize further putative kinases discovered in the future that do not have a defined function. This project is of importance because proteins are biomolecules responsible for as organisms' survival and understanding their unique function is essential for advances in modern medicine, scientific research, and agriculture. |
== Conclusions/Future Direction == | == Conclusions/Future Direction == | ||
Conclusions: | Conclusions: | ||
| - | - D-Glucose exhibited a specific activity of | + | - Authenticated and verified protein function using kinase characterization protocols |
| + | |||
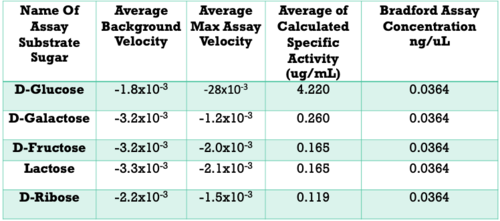
| + | - D-Glucose exhibited a specific activity of ___ ug/mL | ||
- Specific activity: D-Glucose >> D-Galactose > D-Fructose > Lactose > D-Ribose | - Specific activity: D-Glucose >> D-Galactose > D-Fructose > Lactose > D-Ribose | ||
| Line 40: | Line 42: | ||
Future Direction: | Future Direction: | ||
| - | |||
| - | - Verify experimental results | ||
- Publish results for further application | - Publish results for further application | ||
| - | - Authenticate protein function using further kinase characterization protocols | ||
Revision as of 16:24, 21 September 2022
Characterization of the 3r8e Protein, a Novel Glucose Kinase
| |||||||||||
References
1. Blastp [Internet]. Bethesda (MD): Natiobal Library of Medicine (US), National Center for Biotechnology Information; 2004- [cited 2022 March]. Available from: (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins)
2. BASIL. https://basilbiochem.github.io/basil/
3. Holm L (2020) Using Dali for protein structure comparison. Methods Mol. Biol. 2112, 29-42.
4. Small- Molecule Library Screening by Docking with PyRx. .Dallakyan S, Olson AJ Methods Mol Biol. 2015;1263:243-50. The full-text is available at https://www.researchgate.net/publications/2739554875. Small-Molecule Library Screening by Docking with PyRx.
5. Pfam: The Protein families database in 2021 J. Mistry, S. Chuguransky, L. Williams, M. Qureshi, G.A. Salazar, E.L.L. Sonnhammer, S.C.E. Tosatto, L. Paladin, S. Raj, L.J. Richardson, R.D. Finn, A. Bateman Nucleic Acids Research (2020) doi: 10.1093/nar/gkaa913
6. The PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC.
Proteopedia Page Contributors and Editors (what is this?)
Dalton Dencklau, Michel Evertsen, Bonnie Hall, Jaime Prilusky