This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Matheus Andrade Bettiol/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
RhoA (Ras homology gene family member A) is a protein of the small GTPase family. It can be in two conformations, <scene name='97/973102/Rhoa_gtp/1'>linked to GTP</scene> and therefore active, or <scene name='97/973102/Rhoa_gdp/4'>linked to GDP</scene> and consequently inactive. Three factors regulate these two states: | RhoA (Ras homology gene family member A) is a protein of the small GTPase family. It can be in two conformations, <scene name='97/973102/Rhoa_gtp/1'>linked to GTP</scene> and therefore active, or <scene name='97/973102/Rhoa_gdp/4'>linked to GDP</scene> and consequently inactive. Three factors regulate these two states: | ||
| - | 1. GEF (Guanine nucleotide exchange factors): promotes the exchange of GDP for GTP, activating RhoA | + | '''1.''' GEF (Guanine nucleotide exchange factors): promotes the exchange of GDP for GTP, activating RhoA |
| - | 2. GAP (GTPase activating proteins): accelerates the hydrolysis of GTP, inhibiting RhoA | + | '''2.''' GAP (GTPase activating proteins): accelerates the hydrolysis of GTP, inhibiting RhoA |
| - | 3. GDI ([[Guanine nucleotide dissociation inhibitor]]): translocates the membrane GTPase, sequestering it to the cytosol, also inhibiting RhoA | + | '''3.''' GDI ([[Guanine nucleotide dissociation inhibitor]]): translocates the membrane GTPase, sequestering it to the cytosol, also inhibiting RhoA |
| Line 48: | Line 48: | ||
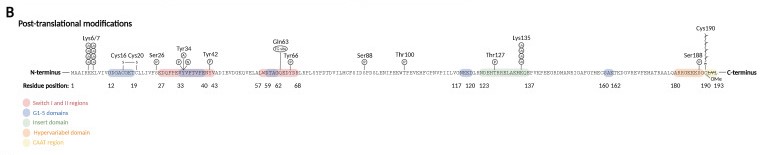
== Post-Translational Modifications == | == Post-Translational Modifications == | ||
| - | Prenylation: the activation of [[Rho GTPase]] require membrane binding, which is necessary for the interaction with membranous GEFs. The membrane association requires C-terminal prenylation, which involves the addition of a geranylgeranyl (20-carbon chain) to Cys190 in the CAAX motif. | + | '''Prenylation:''' the activation of [[Rho GTPase]] require membrane binding, which is necessary for the interaction with membranous GEFs. The membrane association requires C-terminal prenylation, which involves the addition of a geranylgeranyl (20-carbon chain) to Cys190 in the CAAX motif. |
| - | Phosphorylation: can alter the subcellular localization of RhoA when occurs close to C-terminal lipid modifications. On the other hand, phosphorylation of the G-domain affects GTP/GDP cycling and the interaction with effector proteins. | + | '''Phosphorylation:''' can alter the subcellular localization of RhoA when occurs close to C-terminal lipid modifications. On the other hand, phosphorylation of the G-domain affects GTP/GDP cycling and the interaction with effector proteins. |
| - | Oxidation: RhoA can be oxidized on Cys16 and Cys20 (G1 domain), generating a disulfide bond that prevents guanine binding and GEF association, inactivating RhoA. However, if Tyr42 is phosphorylated, serving as a binding site for GEF, oxidation on Cys16/20 reduces the affinity of RhoA for GDI and increases the association with GEF, leading to RhoA activation. | + | '''Oxidation:''' RhoA can be oxidized on Cys16 and Cys20 (G1 domain), generating a disulfide bond that prevents guanine binding and GEF association, inactivating RhoA. However, if Tyr42 is phosphorylated, serving as a binding site for GEF, oxidation on Cys16/20 reduces the affinity of RhoA for GDI and increases the association with GEF, leading to RhoA activation. |
| - | Nitration: nitration on RhoA's Tyr34 (switch I region) introduces a negative charge that modifies the protein structure and leads to a faster GDP release and GTP reload, increasing RhoA activity. | + | '''Nitration:''' nitration on RhoA's Tyr34 (switch I region) introduces a negative charge that modifies the protein structure and leads to a faster GDP release and GTP reload, increasing RhoA activity. |
| - | Adenylation: adenylation on Tyr34 (switch I region) leads to RhoA inhibition. | + | '''Adenylation:''' adenylation on Tyr34 (switch I region) leads to RhoA inhibition. |
Ubiquitination: target the protein for degradation by the proteasome. RhoA is ubiquitylated by E3 [[ubiquitin protein ligase]] complexes, that ubiquitinate either active RhoA on Lys6 and Lys7, inactive RhoA, or both states on Lys135. | Ubiquitination: target the protein for degradation by the proteasome. RhoA is ubiquitylated by E3 [[ubiquitin protein ligase]] complexes, that ubiquitinate either active RhoA on Lys6 and Lys7, inactive RhoA, or both states on Lys135. | ||
[[Image:Pos-translational_modifications_RhoA.jpg]] | [[Image:Pos-translational_modifications_RhoA.jpg]] | ||
| - | Image from | + | Image from Schmidt SI, Blaabjerg M, Freude K, Meyer M. RhoA Signaling in Neurodegenerative Diseases. Cells. 2022 May 1;11(9):1520. |
This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">Secondary Structure</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">Secondary Structure</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
Revision as of 21:30, 25 June 2023
==rhoA==
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644