This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Wayne Chang
From Proteopedia
(→Script Exercises) |
|||
| Line 15: | Line 15: | ||
<applet load='2qc8' size='300' frame='true' align='right' caption='Script Exercises' /> | <applet load='2qc8' size='300' frame='true' align='right' caption='Script Exercises' /> | ||
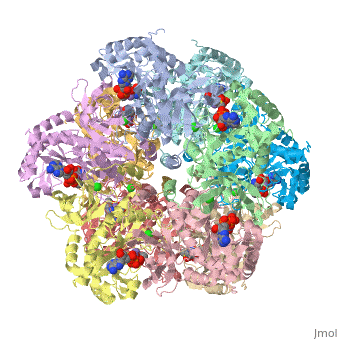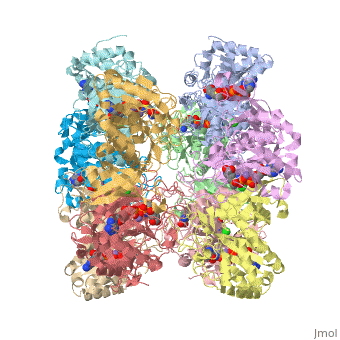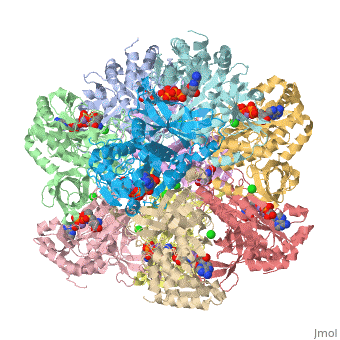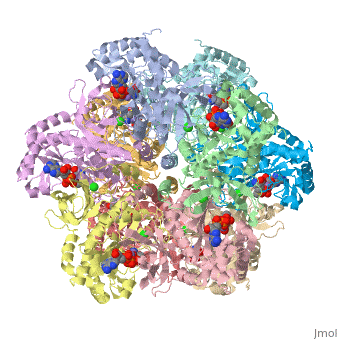
<scene name='User:Wayne_Chang/Glutamate_synthase/1'>Exercise 1: Backbone Trace with ligand</scene> | <scene name='User:Wayne_Chang/Glutamate_synthase/1'>Exercise 1: Backbone Trace with ligand</scene> | ||
| + | Exercise shows a backbone trace of Glutamate Synthase which allows the ligands inside ADP, P3S, Cl- and Mn2+ to be seen. | ||
<scene name='User:Wayne_Chang/Glutamate_synthase/2'>Exercise 2: Ligand and Chain Selection with Labeling</scene> | <scene name='User:Wayne_Chang/Glutamate_synthase/2'>Exercise 2: Ligand and Chain Selection with Labeling</scene> | ||
| + | Isolates chain A of Glutamate Synthase and labels the ligands for easy identification. | ||
<scene name='User:Wayne_Chang/Glutamate_synthase/3'>Exercise 3: Active Site Residues</scene> | <scene name='User:Wayne_Chang/Glutamate_synthase/3'>Exercise 3: Active Site Residues</scene> | ||
| + | Wire Structure of Active Site residues of chain A using information obtained from PDBsum entry for Glutamate Synthase. | ||
<scene name='User:Wayne_Chang/Glutamate_synthase/4'>Exercise 4: Going Solo</scene> | <scene name='User:Wayne_Chang/Glutamate_synthase/4'>Exercise 4: Going Solo</scene> | ||
| - | + | Still picture of salt bridge between residue 63 of chain F and residue 319 of chain G. Bridge length is also provided in angstroms. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
== Outline == | == Outline == | ||
Revision as of 22:02, 15 November 2008
Contents |
Assignment 12: IVC: Ammonium Binding Site
Mapping the Ammonium binding site and explaining how it contributes to catalysis.
Chang, Wayne and Kaushal, Pankaj.
University of Maryland, Baltimore County (UMBC).
Script Exercises
|
Exercise shows a backbone trace of Glutamate Synthase which allows the ligands inside ADP, P3S, Cl- and Mn2+ to be seen.
Isolates chain A of Glutamate Synthase and labels the ligands for easy identification.
Wire Structure of Active Site residues of chain A using information obtained from PDBsum entry for Glutamate Synthase.
Still picture of salt bridge between residue 63 of chain F and residue 319 of chain G. Bridge length is also provided in angstroms.
Outline
-- Work in Progress --
Glutamine synthetase (GS) catalyzes the ATP dependent condensation of glutamate and ammonia, producing, glutamine, ADP, and an inorganic phosphate group.
Glutamate + ATP + NH3 → Glutamine + ADP + phosphate
Ammonium ion is thought to bind to GS at the monovalent cation binding site for Tl(+) and Cs(+) ions.
References
1. Liaw, S-H, et.al.,Discovery of the ammonium substrate site on glutamine synthetase, a third cation binding site Protein Sci. 1995 4: 2358-2365[1]