From Proteopedia
(Difference between revisions)
proteopedia linkproteopedia link
|
|
| Line 48: |
Line 48: |
| | | | |
| | | | |
| - | <Structure load='2BBM' size='300' frame='true' align='RIGHT' caption='Calmodulin bound to a target peptide' scene='Insert optional scene name here' /> | + | <Structure load='2BBM' size='300' frame='true' align='RIGHT' caption='Calmodulin bound to a target peptide' scene=''''Ca<sup>2+</sup>-bound calmodulin'Ca<sup>2+</sup>-bound calmodulin'/> |
| | *'''Calmodulin bound to a target peptide''' | | *'''Calmodulin bound to a target peptide''' |
| | | | |
Revision as of 19:59, 30 December 2011
Calmodulin (CaM) for Calcium-Modulated protein is an important protein that intervenes in a wide range of activities inflammation, metabolism, apoptosis, smooth muscle contraction, intracellular movement, short-term and long-term memory, and the immune response.
Indeed,it is a small (16.7 kDa = 148 aa) and highly conserved protein that is necessary in all eukaryotic cells because it represents an essential calcium sensor with troponin C its isoform.
Calmodulin contains four Ca2+ binding sites and the binding of calcium induces a conformational change in calmodulin that can cause the activation of key enzymes such as kinases or phosphatases proteins (especially phosphorylase kinases) which are not necessarily themselves Ca2+-sensitive and allows a large diversity of cellular response.
Structure
Calmodulin [1]
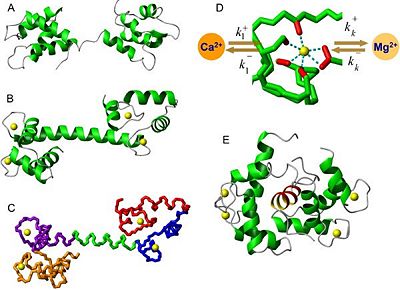
The calmodulin structure has been determined by NMR. This method reveals that calmodulin is a long molecule which looks like a dumbbell because it contains two globular domains (the and the ) linked by a .
Each lobe contains a pair of motifs (called EF-hand) that can bind two Ca2+ ions. However those lobes do not have the same properties because the C-lobe has higher Ca2+ affinity than the N-lobe.
The two EF-hands are located in the vicinity of each other. Those neighboring sited are very likely to structurally influence each other upon Ca2+ binding to one of them.
The affinity of the individual Ca2+ ion binding sites are in the range 10−5-10−6 mol.L−1 and adjacent sites bind Ca2+ with positive cooperativity, so that attachment of the first Ca2+ ion enhances the affinity of its neighbour. This has the effect of making the protein sensitive to small changes in the concentration of Ca2+ within the signaling range.
Ca2+-calmodulin itself has no intrinsic catalytic activity. Its action depends on its close association with a target enzyme.
| |
- Three-dimensional structure of apocalmodulin
In the absence of bound Ca2+, the helices of calmodulin pack so that their hydrophobic side chains are not exposed. In this form it is unable to interact with its targets[1].
Binding of Ca2+ to the four sites induces a large conformational change causing the terminal helices to expose hydrophobic surfaces and also a long central α-helical segment. Ca2+-bound calmodulin binds to its targets with high affinity (KD≈10−9 mol.L−1).
| 2+</sup>-bound calmodulin'Ca2+-bound calmodulin'/>
- Calmodulin bound to a target peptide
To form the bound state, the central residues of the link region unwind form their α-helical arrangement to form a hinge that allows the molecule to bend and wrap itself around the target.
The N-terminal and C-terminal regions approach each other and by their hydrophobic surfaces bind to it, rather like two hands holding a rope.
This encourages the target sequence to adopt an α-helical arrangement so that it occupies the center of a hydrophobic tunnel.
The consequence of this interaction is a conformational change in the target, a state that persists only as long as the Ca2+ concentration remains high.
When the Ca2+ concentration falls, calcium dissociates and calmodulin is quickly released, inactivating the target. However, at least one important target protein is an exception to this rule. This is CaM-kinase II which can retain its active state after it has been activated by calmodulin.
CaMII kinase
Calmodulin plays an important role through kinase enzymes such as calcium/calmodulin-dependent kinase II (CaMKII) that is a multifunctional serine/threonine kinase found in many tissues. Activation of CaMKII contributes to synaptic plasticity and regulation of excitory synaptic transmission. The regulatory domain of CaMKII contains an autophosphorylation site, which is essential for its calcium-dependent activation.
Family members
- Calmodulin 1 (CALM1)
- Calmodulin 2 (CALM2)
- Calmodulin 3 (CALM3)
- Calmodulin-like 1 (CALML1)
- Calmodulin-like 3 (CALML3)
- Calmodulin-like 4 (CALML4)
- Calmodulin-like 5 (CALML5)
- Calmodulin-like 6 (CALML6)
External Resources
References
- Valeyev NV, Bates DG, Heslop-Harrison P, Postlethwaite I, Kotov NV. Elucidating the mechanisms of cooperative calcium-calmodulin interactions: a structural systems biology approach. BMC Syst Biol. 2008 Jun 2;2:48. PMID:18518982 doi:10.1186/1752-0509-2-48
- Fallon JL, Quiocho FA. A closed compact structure of native Ca(2+)-calmodulin. Structure. 2003 Oct;11(10):1303-7. PMID:14527397
- Evans TI, Shea MA. Energetics of calmodulin domain interactions with the calmodulin binding domain of CaMKII. Proteins. 2009 Jul;76(1):47-61. PMID:19089983 doi:10.1002/prot.22317
- Halling DB, Georgiou DK, Black DJ, Yang G, Fallon JL, Quiocho FA, Pedersen SE, Hamilton SL. Determinants in CaV1 channels that regulate the Ca2+ sensitivity of bound calmodulin. J Biol Chem. 2009 Jul 24;284(30):20041-51. Epub 2009 May 27. PMID:19473981 doi:10.1074/jbc.M109.013326
- Colbran RJ, Brown AM. Calcium/calmodulin-dependent protein kinase II and synaptic plasticity. Curr Opin Neurobiol. 2004 Jun;14(3):318-27. PMID:15194112 doi:10.1016/j.conb.2004.05.008
Proteopedia page contributors and editor
Tonazzini Saphia, Planchenault Charlène
|