This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox 645
From Proteopedia
(Difference between revisions)
| Line 16: | Line 16: | ||
In the mid 1980's, the structure of HIV-1 Protease was hypothesized by Lawrence Pearl and William Taylor to consist of a single domain of the eukaryotic aspartic protease and function in the dimeric form. As NMR was not yet in use at this time, x-ray crystallography was implored but required a large quantity of large crystals. This was problematic as heavy atom derivatives had to be used to overcome the phase problems with this new structure and a considerable amount of protein was needed. Eventually these problems were overcome and the first images of HIV-1 protease where announced by the Merck group, who were most successful at large-scale protein purification and crystallization. | In the mid 1980's, the structure of HIV-1 Protease was hypothesized by Lawrence Pearl and William Taylor to consist of a single domain of the eukaryotic aspartic protease and function in the dimeric form. As NMR was not yet in use at this time, x-ray crystallography was implored but required a large quantity of large crystals. This was problematic as heavy atom derivatives had to be used to overcome the phase problems with this new structure and a considerable amount of protein was needed. Eventually these problems were overcome and the first images of HIV-1 protease where announced by the Merck group, who were most successful at large-scale protein purification and crystallization. | ||
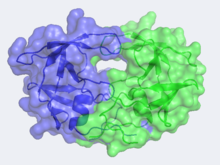
| - | Unlike most members of the aspartyl protease class, which generally exist as two domain monomers, HIV protease is a dimmer with two identical <scene name='Sandbox_645/Monomer/2'>subunits</scene> that are comprised of 99 amino acids. These two subunits come together to form a very narrow 7.46 angstrom diameter<scene name='User:David_Canner/Sandbox_HIV/Narrow_Tunnel/1'>tunnel</scene> for the ligand to navigate; enclosing the active site in the middle. Instead of one flap (a long flexible β-hairpin loop from the N-terminal domain) closing over the active site like in pepsins, the homodimeric retroviral enzyme has two poorly ordered flaps.<ref>PMID:20593466</ref>The polyprotein is capable of making it to the active site not by squeezing through the narrow tunnel but rather by having the flaps open. Each subunit donates the catalytic triad, Asp-Ser-Gly, to the highly conserved active site, which also closely resembles that of monomeric proteases like pepsin. | + | Unlike most members of the aspartyl protease class, which generally exist as two domain monomers, HIV protease is a dimmer with two identical <scene name='Sandbox_645/Monomer/2'>subunits</scene> that are comprised of 99 amino acids. These two subunits come together to form a very narrow 7.46 angstrom diameter <scene name='User:David_Canner/Sandbox_HIV/Narrow_Tunnel/1'>tunnel</scene> for the ligand to navigate; enclosing the active site in the middle. Instead of one flap (a long flexible β-hairpin loop from the N-terminal domain) closing over the active site like in pepsins, the homodimeric retroviral enzyme has two poorly ordered flaps.<ref>PMID:20593466</ref>The polyprotein is capable of making it to the active site not by squeezing through the narrow tunnel but rather by having the flaps <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph/3'>open</scene>. Each subunit donates the catalytic triad, Asp-Ser-Gly, to the highly conserved active site, which also closely resembles that of monomeric proteases like pepsin. |
The X-ray structure of HIV-1 protease reveals that it is composed of <scene name='User:David_Canner/Sandbox_HIV/Identical_subunits/1'>two symmetrically related subunits</scene>, each consisting of 99 amino acid residues. The subunits come together in such as way as to form a tunnel where they meet. This tunnel is of critical importance because the active site of the protease is located in its interior. The active site consists of <scene name='User:David_Canner/Sandbox_HIV/Catalytic_triad/3'> two Asp-Thr-Gly conserved sequences</scene>, making it a member of the aspartyl protease family. The two Asp's are <scene name='User:David_Canner/Sandbox_HIV/Catalytic_asp/1'>essential catalytic residues</scene> that activate a water molecule to hydrolytically cleave the polyprotein that binds in the tunnel.<ref>PMID:1799632</ref> You may be wondering how a polyprotein makes its way into the active-site tunnel, as the<scene name='User:David_Canner/Sandbox_HIV/Narrow_tunnel/1'> tunnel appears to be too narrow </scene> to admit it. The key is the two flexible flaps on the top of the tunnel that <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph/3'>move to allow proteins </scene>to enter the tunnel. The flaps <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph_flaps/2'>undergo a dramatic movement</scene>, shifting from an open to a closed conformation to bind the target in an appropriate conformation for cleavage. | The X-ray structure of HIV-1 protease reveals that it is composed of <scene name='User:David_Canner/Sandbox_HIV/Identical_subunits/1'>two symmetrically related subunits</scene>, each consisting of 99 amino acid residues. The subunits come together in such as way as to form a tunnel where they meet. This tunnel is of critical importance because the active site of the protease is located in its interior. The active site consists of <scene name='User:David_Canner/Sandbox_HIV/Catalytic_triad/3'> two Asp-Thr-Gly conserved sequences</scene>, making it a member of the aspartyl protease family. The two Asp's are <scene name='User:David_Canner/Sandbox_HIV/Catalytic_asp/1'>essential catalytic residues</scene> that activate a water molecule to hydrolytically cleave the polyprotein that binds in the tunnel.<ref>PMID:1799632</ref> You may be wondering how a polyprotein makes its way into the active-site tunnel, as the<scene name='User:David_Canner/Sandbox_HIV/Narrow_tunnel/1'> tunnel appears to be too narrow </scene> to admit it. The key is the two flexible flaps on the top of the tunnel that <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph/3'>move to allow proteins </scene>to enter the tunnel. The flaps <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph_flaps/2'>undergo a dramatic movement</scene>, shifting from an open to a closed conformation to bind the target in an appropriate conformation for cleavage. | ||
Revision as of 08:15, 27 November 2012
HIV-1 Protease
| |||||||||||