From Proteopedia
(Difference between revisions)
proteopedia linkproteopedia link
|
|
| Line 33: |
Line 33: |
| | | | |
| | == Clinical relevence == | | == Clinical relevence == |
| | + | {| align=middle |
| | + | |- |
| | + | | |
| | + | |
| | + | [[Image:picture 3.jpg | 300 px | thumb |Figure 1 showing the methylation of the histone protein and the subsequent formation of heterochromatin]] |
| | + | |
| | ---- | | ---- |
| | | | |
| | == References == | | == References == |
| | -- | | -- |
Revision as of 20:27, 19 April 2013
==Your Heading Here (maybe something like 'Structure'-- PLEASE DO NOT DELETE THIS TEMPLATE -->
| This Sandbox is Reserved from Feb 1, 2013, through May 10, 2013 for use in the course "Biochemistry" taught by Irma Santoro at the Reinhardt University. This reservation includes Sandbox Reserved 591 through Sandbox Reserved 599.
|
To get started:
- Click the edit this page tab at the top. Save the page after each step, then edit it again.
- Click the 3D button (when editing, above the wikitext box) to insert Jmol.
- show the Scene authoring tools, create a molecular scene, and save it. Copy the green link into the page.
- Add a description of your scene. Use the buttons above the wikitext box for bold, italics, links, headlines, etc.
More help: Help:Editing
|
Background
SUV39h1 is part of a class of enzymes that deals with the methyltransferase of histone proteins in nucleosomes. The methylation of histone proteins is an essential part of an area of genetics which deals with the modification of histone proteins called epigenomics. Epigenomics is the study of modifications of nucleosomes, which inhibit or express gene trancriscription without changing the underlining DNA sequence. These changes are allowed because the amino acid tails which extend out and away from the histone porteins. Methylation of histone protein is one of many ways that regulates gene expression. The regulation of gene expression is dependent on the state of the gene. A gene cannot be transcribed when the promoter of the gene is wrapped around the nucleosome, forming a heterochromatin state; thus, the gene is inhibited from being transcribed. coversly Methylation of the histone protein causes the winding around the nucleosomes, transforming a euchromatin state into a heterochromatin state. Of course, SUV39H1 alone does not methtylate the histone proteins. SUV39H1 interacts with other proteins in order to efficiently methylate a histone protein (shown in Figure 1).
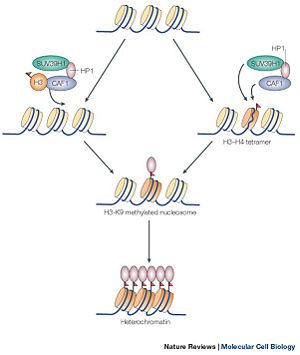 Figure 1 showing the methylation of the histone protein and the subsequent formation of heterochromatin
Structure
Crystal structure shows two main parts to the protein. There is a chromodomain and SET domain. The chromodomain starts at the N-terminus of the enzyme and continues toward the C-terminus, where the SET catalytic domain is located. The chromodomain length is around 44-106 amino acids long, which forms three antiparellel beta sheets. The lengths for each of three beta sheets are 45-53, 58-64 and 73-76 amino acids long for beta 1, beta 2 and beta 3, respectively. These three beta sheets form the the chromodomain
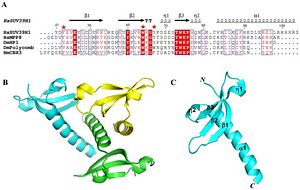 Figure 2 shows the residue and tertiary and Quaternary structure SUV39H1
Function
There are two types of histone methyltransferases: lysine specific and argine specific; each are residues which the enzyme transfers a mathyl group to. Within the lysine specific, there are further two main types: SET (Su(var)3-9, Enhancer of Zeste, Trithorax) and non-SET domains. These domains are an essential part of the enzyme because the main catalytic occurs in SET domain of the methyltransferase. SUV39h1 is considered to have a lysine specific SET domain which catalyzes the methyltransferase. In the nucleosomes, there are four different proteins; each consists of two copies which make the entire nucleosome. The mechanism by which SUV39H1 methylates the histone proteins is also known. A nearby tyrosine residue in the histone protein creates a strong nucleophile by deprotonating a nearby lysine residue in the same histone protein. The lysine residue is a very strong nucleophile which attacks the sulfur atom of the cofactor S-Adensyl methoinine (SAM)and extracts a methyl group from the cofactor. subsequently, the attack methylates the histone protein. The cofactor is very important because it is the source of the methyl group.
---
Clinical relevence
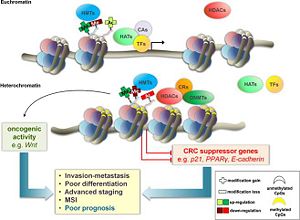 Figure 1 showing the methylation of the histone protein and the subsequent formation of heterochromatin
References
--
|
|
|



