This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 967
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
== Biological role == | == Biological role == | ||
Ribonucleases H are the only known enzymes, able to degrade the RNA strand of a DNA/RNA hybrid in a sequence-nonspecific way. | Ribonucleases H are the only known enzymes, able to degrade the RNA strand of a DNA/RNA hybrid in a sequence-nonspecific way. | ||
| - | There are two types of RNase H (RNases H1 and RNases H2) classified according to their sequence conservation and substrate preference. Currently, three types of RNA/DNA hybrids are known: simple RNA/DNA duplexes ('''Figure 1A'''), RNA•DNA/DNA hybrids ('''Figure 1B'''), and DNA•RNAfew•DNA/DNA hybrids ('''Figure 1C'''). RNases H2 is totally able to cleave a single ribonucleotide embedded in a double strand DNA (DNA• RNAfew •DNA/DNA type) when RNases H1 require at least 4 ribonucleotides. This ability and their high expression in proliferating cells suggest that RNases H2 are involved in DNA repair and replication<ref> Rychlik, Monika P., Hyongi Chon, Susana M. Cerritelli, Paulina Klimek, Robert J. Crouch, and Marcin Nowotny. “Crystal Structures of RNase H2 in Complex with Nucleic Acid Reveal the Mechanism of RNA-DNA Junction Recognition and Cleavage.” Molecular Cell 40, no. 4 (November 24, 2010): 658–70. | + | There are two types of RNase H (RNases H1 and RNases H2) classified according to their sequence conservation and substrate preference. Currently, three types of RNA/DNA hybrids are known: simple RNA/DNA duplexes ('''Figure 1A'''), RNA•DNA/DNA hybrids ('''Figure 1B'''), and DNA•RNAfew•DNA/DNA hybrids ('''Figure 1C'''). RNases H2 is totally able to cleave a single ribonucleotide embedded in a double strand DNA (DNA• RNAfew •DNA/DNA type) when RNases H1 require at least 4 ribonucleotides. This ability and their high expression in proliferating cells suggest that RNases H2 are involved in DNA repair and replication<ref> Rychlik, Monika P., Hyongi Chon, Susana M. Cerritelli, Paulina Klimek, Robert J. Crouch, and Marcin Nowotny. “Crystal Structures of RNase H2 in Complex with Nucleic Acid Reveal the Mechanism of RNA-DNA Junction Recognition and Cleavage.” Molecular Cell 40, no. 4 (November 24, 2010): 658–70. doi:10.1016/j.molcel.2010.11.001.</ref>. |
[[Image:ThreetypesofRNA-DNAhybrids.png|300px|left|thumb| '''Figure 1''' : Three types of RNA/DNA hybrids]] | [[Image:ThreetypesofRNA-DNAhybrids.png|300px|left|thumb| '''Figure 1''' : Three types of RNA/DNA hybrids]] | ||
| - | Indeed, ribonucleotides are wrongly incorporated into DNA during DNA replication at a frequency of about 2 ribonucleotides per kb. With such frequency, these errors are by far the most abundant threat of DNA damaging. Hence, a correction is essential to the preservation of DNA integrity: the most common correction mechanism involves RNases H2 and is called Ribonucleotide Excision Repair (RER). The incorporation of ribonucleotides in DNA produce DNA•RNAfew•DNA/DNA hybrids from which the few misincorporated ribonucleotides can be removed by an RNase H2<ref> Sparks, Justin L., Hyongi Chon, Susana M. Cerritelli, Thomas A. Kunkel, Erik Johansson, Robert J. Crouch, and Peter M. Burgers. “RNase H2-Initiated Ribonucleotide Excision Repair.” Molecular Cell 47, no. 6 (September 28, 2012): 980–86. | + | Indeed, ribonucleotides are wrongly incorporated into DNA during DNA replication at a frequency of about 2 ribonucleotides per kb. With such frequency, these errors are by far the most abundant threat of DNA damaging. Hence, a correction is essential to the preservation of DNA integrity: the most common correction mechanism involves RNases H2 and is called Ribonucleotide Excision Repair (RER). The incorporation of ribonucleotides in DNA produce DNA•RNAfew•DNA/DNA hybrids from which the few misincorporated ribonucleotides can be removed by an RNase H2<ref> Sparks, Justin L., Hyongi Chon, Susana M. Cerritelli, Thomas A. Kunkel, Erik Johansson, Robert J. Crouch, and Peter M. Burgers. “RNase H2-Initiated Ribonucleotide Excision Repair.” Molecular Cell 47, no. 6 (September 28, 2012): 980–86. doi:10.1016/j.molcel.2012.06.035.</ref>. |
| - | This repair activity is guided by the interaction between C-terminus of RNase H2B protein and the DNA clamp PCNA. This interaction occurs through a hydrophobic conserved peptide motif called the PCNA interaction peptide PIP (PIP-box: Residues 294 to 301 MKSIDTFF of H2B protein) that interacts with a hydrophobic groove near the PCNA C-terminus. This interaction allows RNase H2 to scan DNA for misincorporated ribonucleotides which makes the Ribonucleotide Excision Repair more efficient<ref | + | This repair activity is guided by the interaction between C-terminus of RNase H2B protein and the DNA clamp PCNA. This interaction occurs through a hydrophobic conserved peptide motif called the PCNA interaction peptide PIP (PIP-box: Residues 294 to 301 MKSIDTFF of H2B protein) that interacts with a hydrophobic groove near the PCNA C-terminus. This interaction allows RNase H2 to scan DNA for misincorporated ribonucleotides which makes the Ribonucleotide Excision Repair more efficient<ref> Bubeck, Doryen, Martin A. M. Reijns, Stephen C. Graham, Katy R. Astell, E. Yvonne Jones, and Andrew P. Jackson. “PCNA Directs Type 2 RNase H Activity on DNA Replication and Repair Substrates.” Nucleic Acids Research 39, no. 9 (May 2011): 3652–66. [http://dx.doi.org/10.1093/nar/gkq980 doi:10.1093/nar/gkq980.</ref>. |
| - | Furthermore, ''in vitro'' studies have shown that RNases H2 is likely to be involved in the removal of RNA primer from Okazaki fragment produced during the synthesis of the lagging strand in DNA replication since Okazaki fragment are RNA•DNA/DNA hybrids ('''Figure 1B''') | + | Furthermore, ''in vitro'' studies have shown that RNases H2 is likely to be involved in the removal of RNA primer from Okazaki fragment produced during the synthesis of the lagging strand in DNA replication since Okazaki fragment are RNA•DNA/DNA hybrids ('''Figure 1B'''). |
| - | RNases H2 activity is crucial in mammalian cells, for instance a mutation in human RNase H2 causes Aicardi-Goutières syndrome. This syndrome is an auto-inflammatory disorder that may be the consequence of an increased production of incorrect nucleic acid by-products during DNA replication | + | RNases H2 activity is crucial in mammalian cells, for instance a mutation in human RNase H2 causes Aicardi-Goutières syndrome. This syndrome is an auto-inflammatory disorder that may be the consequence of an increased production of incorrect nucleic acid by-products during DNA replication. |
| Line 22: | Line 22: | ||
=== A heteromeric complex === | === A heteromeric complex === | ||
| - | It has been shown that the Mammalian RNase complex is a heteromeric complex formed by 3 distinct proteins: <scene name='60/604486/H2a/1'>H2A</scene>,<scene name='60/604486/H2b/1'>H2B</scene> and <scene name='60/604486/H2c/1'>H2C</scene>. H2A protein is the catalytic subunit and H2B/H2C proteins are auxiliary subunits: they are structural domains that facilitate cohesion of the complex | + | It has been shown that the Mammalian RNase complex is a heteromeric complex formed by 3 distinct proteins: <scene name='60/604486/H2a/1'>H2A</scene>,<scene name='60/604486/H2b/1'>H2B</scene> and <scene name='60/604486/H2c/1'>H2C</scene>. H2A protein is the catalytic subunit and H2B/H2C proteins are auxiliary subunits: they are structural domains that facilitate cohesion of the complex. |
The first domain structure of the complex, H2A, contains 301 amino acids, almost as H2B protein which computes 308 amino acids. H2C protein is the smallest subunit: it only has 166 amino acids. | The first domain structure of the complex, H2A, contains 301 amino acids, almost as H2B protein which computes 308 amino acids. H2C protein is the smallest subunit: it only has 166 amino acids. | ||
Each of these proteins adopts various secondary structures with β-strands and α-helices: | Each of these proteins adopts various secondary structures with β-strands and α-helices: | ||
| - | * H2A protein has 12 α-helices, 11 β-strands and 3 turns | + | * H2A protein has 12 α-helices, 11 β-strands and 3 turns, |
| - | * H2B molecule computes 8 α-helices, 7 β-strands and 3 turns | + | * H2B molecule computes 8 α-helices, 7 β-strands and 3 turns, |
| - | * H2C subunit consists of 5 α-helices, 8 β-strands and 2 turns | + | * H2C subunit consists of 5 α-helices, 8 β-strands and 2 turns. |
=== Several interactions between the subunits === | === Several interactions between the subunits === | ||
H2C protein is found in the middle of the elongated complex structure, flanked by H2A and H2B proteins on the ends. | H2C protein is found in the middle of the elongated complex structure, flanked by H2A and H2B proteins on the ends. | ||
| - | The complex is stabilized by the intimately interwoven architecture of H2B and H2C: The N-terminal region of H2B protein (amino acids 1-92) weaves together with H2C domain to form 3 β-barrels, also called “triple barrel”. This triple barrel is formed from a total of 18 β-sheets and produces a pseudo-2-fold axis of symmetry along the central barrel. Also, it permits to leave the mostly α-helical C-terminal region of H2B available for potential interactions with other protein (for example the PCNA protein). Finally, it has been found that the motif provides a platform for securely binding the H2A protein: the side and end of the first barrel in the subcomplex H2B/H2C form a tight interface with amino acids 197-258 in the C-terminal region of H2A protein. This interface is composed mainly of hydrophobic residues. | + | The complex is stabilized by the intimately interwoven architecture of H2B and H2C: The N-terminal region of H2B protein (amino acids 1-92) weaves together with H2C domain to form 3 β-barrels, also called <scene name='60/604486/Triple_barrel/2'>“triple barrel”</scene>. This triple barrel is formed from a total of <scene name='60/604486/Triple_barrel/1'>18 β-sheets</scene> and produces a pseudo-2-fold axis of symmetry along the central barrel. Also, it permits to leave the mostly α-helical C-terminal region of H2B available for potential interactions with other protein (for example the PCNA protein). Finally, it has been found that the motif provides a platform for securely binding the H2A protein: the side and end of the first barrel in the subcomplex H2B/H2C form <scene name='60/604486/Tight_interface_h2ah2c/1'> a tight interface </scene> with amino acids 197-258 in the C-terminal region of H2A protein. This interface is composed mainly of hydrophobic residues. |
| Line 39: | Line 39: | ||
It has been proved that the position of RNA/DNA complex in the active site cleft is determined by several favorable electrostatic interactions between the nucleic acid and positively charged amino acids of the protein. | It has been proved that the position of RNA/DNA complex in the active site cleft is determined by several favorable electrostatic interactions between the nucleic acid and positively charged amino acids of the protein. | ||
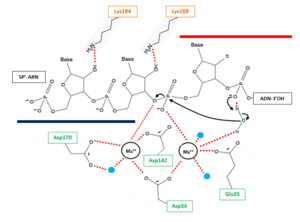
| - | The β6-α6 loop of the H2A protein could play a role in substrate recognition: the minor groove of the double helix molecule straddles this area of the protein, which results in a non-sequence specific cleavage by the enzyme. Moreover, the β6-α6 loop contains a Lysine amino acid in position 128, which might act as a sensor for the hybrid by forming an interaction with the 2’-hydroxyl group of the ribose in the 3’ nucleotide of the RNA primer in the RNA-DNA hybrid ('''Figure 2'''). Therefore, since DNA does not contain a 2’-hydroxyl group in it nucleotide sequence, the RNase H2 can only recognize RNA in the hybrid: only ribonucleotides of the RNA strand are positioned in the active site. The RNA-DNA hybrid is placed such that the target phosphodiester bond between the RNA and DNA parts of the hybrid is in the proper orientation for nucleophile attack by a two-metal ion mechanism. | + | The β6-α6 loop of the H2A protein could play a role in substrate recognition: the minor groove of the double helix molecule straddles this area of the protein, which results in a non-sequence specific cleavage by the enzyme. Moreover, the β6-α6 loop contains a <scene name='60/604486/Site_actif_dna_recongnition/1'>Lysine amino acid in position 128</scene>, which might act as a sensor for the hybrid by forming an interaction with the 2’-hydroxyl group of the ribose in the 3’ nucleotide of the RNA primer in the RNA-DNA hybrid ('''Figure 2'''). Therefore, since DNA does not contain a 2’-hydroxyl group in it nucleotide sequence, the RNase H2 can only recognize RNA in the hybrid: only ribonucleotides of the RNA strand are positioned in the active site. The RNA-DNA hybrid is placed such that the target phosphodiester bond between the RNA and DNA parts of the hybrid is in the proper orientation for nucleophile attack by a two-metal ion mechanism. |
It is important to notice that the Mammalian RNase H2 contains only one cleft with the active site for substrate binding: RNase H2 may recognize single ribonucleotide within a DNA duplex that have a B-form helical structure, as well as longer RNA in RNA-DNA hybrid which adopts intermediate A/B form structure. Thus, the RNase H2 enzyme needs to bind both conformations to able to fully complete all its roles. | It is important to notice that the Mammalian RNase H2 contains only one cleft with the active site for substrate binding: RNase H2 may recognize single ribonucleotide within a DNA duplex that have a B-form helical structure, as well as longer RNA in RNA-DNA hybrid which adopts intermediate A/B form structure. Thus, the RNase H2 enzyme needs to bind both conformations to able to fully complete all its roles. | ||
Revision as of 17:10, 8 January 2015
| This Sandbox is Reserved from 15/11/2014, through 15/05/2015 for use in the course "Biomolecule" taught by Bruno Kieffer at the Strasbourg University. This reservation includes Sandbox Reserved 951 through Sandbox Reserved 975. |
To get started:
More help: Help:Editing |
Structure of the Mouse RNase H2 Complex
| |||||||||||
References
- ↑ http://genome-euro.ucsc.edu/cgi-bin/hgTracks?clade=mammal&org=Mouse&db=mm10&position=RnaseH2&hgt.positionInput=RnaseH2&hgt.suggestTrack=knownGene&Submit=submit&hgsid=201143152_yP1Xd4bMnHS7DV0d3VcqpDSxzzuQ&pix=1563
- ↑ Rychlik, Monika P., Hyongi Chon, Susana M. Cerritelli, Paulina Klimek, Robert J. Crouch, and Marcin Nowotny. “Crystal Structures of RNase H2 in Complex with Nucleic Acid Reveal the Mechanism of RNA-DNA Junction Recognition and Cleavage.” Molecular Cell 40, no. 4 (November 24, 2010): 658–70. doi:10.1016/j.molcel.2010.11.001.
- ↑ Sparks, Justin L., Hyongi Chon, Susana M. Cerritelli, Thomas A. Kunkel, Erik Johansson, Robert J. Crouch, and Peter M. Burgers. “RNase H2-Initiated Ribonucleotide Excision Repair.” Molecular Cell 47, no. 6 (September 28, 2012): 980–86. doi:10.1016/j.molcel.2012.06.035.
- ↑ Bubeck, Doryen, Martin A. M. Reijns, Stephen C. Graham, Katy R. Astell, E. Yvonne Jones, and Andrew P. Jackson. “PCNA Directs Type 2 RNase H Activity on DNA Replication and Repair Substrates.” Nucleic Acids Research 39, no. 9 (May 2011): 3652–66. [http://dx.doi.org/10.1093/nar/gkq980 doi:10.1093/nar/gkq980.