This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 967
From Proteopedia
(Difference between revisions)
| Line 35: | Line 35: | ||
== Interactions with nucleic acids == | == Interactions with nucleic acids == | ||
| - | It has been proved that the position of RNA/DNA complex in the active site cleft is determined by several favorable electrostatic interactions between the nucleic acid and positively charged amino acids of the protein<ref name = "ref2">. | + | It has been proved that the position of RNA/DNA complex in the active site cleft is determined by several favorable electrostatic interactions between the nucleic acid and positively charged amino acids of the protein<ref name = "ref2" />. |
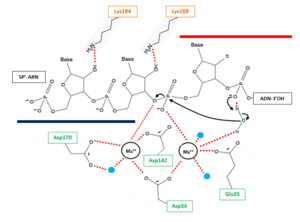
The β6-α6 loop of the H2A protein could play a role in substrate recognition: the minor groove of the double helix molecule straddles this area of the protein, which results in a non-sequence specific cleavage by the enzyme. Moreover, the β6-α6 loop contains a<scene name='60/604486/Site_actif_dna_recongnition/1'>Lysine amino acid in position 128</scene>, which might act as a sensor for the hybrid by forming an interaction with the 2’-hydroxyl group of the ribose in the 3’ nucleotide of the RNA primer in the RNA-DNA hybrid ('''Figure 2'''). Therefore, since DNA does not contain a 2’-hydroxyl group in it nucleotide sequence, the RNase H2 can only recognize RNA in the hybrid: only ribonucleotides of the RNA strand are positioned in the active site. The RNA-DNA hybrid is placed such that the target phosphodiester bond between the RNA and DNA parts of the hybrid is in the proper orientation for nucleophile attack by a two-metal ion mechanism. | The β6-α6 loop of the H2A protein could play a role in substrate recognition: the minor groove of the double helix molecule straddles this area of the protein, which results in a non-sequence specific cleavage by the enzyme. Moreover, the β6-α6 loop contains a<scene name='60/604486/Site_actif_dna_recongnition/1'>Lysine amino acid in position 128</scene>, which might act as a sensor for the hybrid by forming an interaction with the 2’-hydroxyl group of the ribose in the 3’ nucleotide of the RNA primer in the RNA-DNA hybrid ('''Figure 2'''). Therefore, since DNA does not contain a 2’-hydroxyl group in it nucleotide sequence, the RNase H2 can only recognize RNA in the hybrid: only ribonucleotides of the RNA strand are positioned in the active site. The RNA-DNA hybrid is placed such that the target phosphodiester bond between the RNA and DNA parts of the hybrid is in the proper orientation for nucleophile attack by a two-metal ion mechanism. | ||
| Line 56: | Line 56: | ||
== References == | == References == | ||
| - | < | + | <references/> |
== Proteopedia page contributors and editors == | == Proteopedia page contributors and editors == | ||
Anaïs Bourbigot & Valériane Keïta | Anaïs Bourbigot & Valériane Keïta | ||
Revision as of 00:47, 10 January 2015
| This Sandbox is Reserved from 15/11/2014, through 15/05/2015 for use in the course "Biomolecule" taught by Bruno Kieffer at the Strasbourg University. This reservation includes Sandbox Reserved 951 through Sandbox Reserved 975. |
To get started:
More help: Help:Editing |
Structure of the Mouse RNase H2 Complex
| |||||||||||
References
- ↑ http://genome-euro.ucsc.edu/cgi-bin/hgTracks?clade=mammal&org=Mouse&db=mm10&position=RnaseH2&hgt.positionInput=RnaseH2&hgt.suggestTrack=knownGene&Submit=submit&hgsid=201143152_yP1Xd4bMnHS7DV0d3VcqpDSxzzuQ&pix=1563
- ↑ 2.0 2.1 Rychlik, Monika P., Hyongi Chon, Susana M. Cerritelli, Paulina Klimek, Robert J. Crouch, and Marcin Nowotny. “Crystal Structures of RNase H2 in Complex with Nucleic Acid Reveal the Mechanism of RNA-DNA Junction Recognition and Cleavage.” Molecular Cell 40, no. 4 (November 24, 2010): 658–70. doi:10.1016/j.molcel.2010.11.001.
- ↑ Sparks, Justin L., Hyongi Chon, Susana M. Cerritelli, Thomas A. Kunkel, Erik Johansson, Robert J. Crouch, and Peter M. Burgers. “RNase H2-Initiated Ribonucleotide Excision Repair.” Molecular Cell 47, no. 6 (September 28, 2012): 980–86. doi:10.1016/j.molcel.2012.06.035.
- ↑ 4.0 4.1 4.2 Bubeck, Doryen, Martin A. M. Reijns, Stephen C. Graham, Katy R. Astell, E. Yvonne Jones, and Andrew P. Jackson. “PCNA Directs Type 2 RNase H Activity on DNA Replication and Repair Substrates.” Nucleic Acids Research 39, no. 9 (May 2011): 3652–66. doi:10.1093/nar/gkq980.
- ↑ Shaban, Nadine M., Scott Harvey, Fred W. Perrino, and Thomas Hollis. “The Structure of the Mammalian RNase H2 Complex Provides Insight into RNA•DNA Hybrid Processing to Prevent Immune Dysfunction.” Journal of Biological Chemistry 285, no. 6 (February 5, 2010): 3617–24. doi:10.1074/jbc.M109.059048.
- ↑ http://www.uniprot.org/uniprot/Q9CWY8
- ↑ http://www.uniprot.org/uniprot/Q80ZV0
- ↑ http://www.uniprot.org/uniprot/Q9CQ18
- ↑ Nicholson, Allen W. Ribonucleases. Springer Science & Business Media, 2011.
Proteopedia page contributors and editors
Anaïs Bourbigot & Valériane Keïta