This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Amyloid precursor protein
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
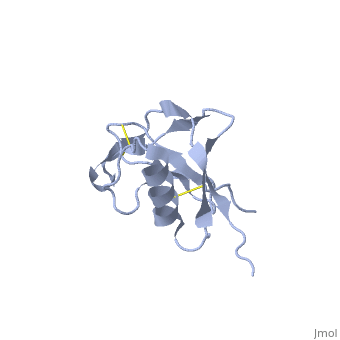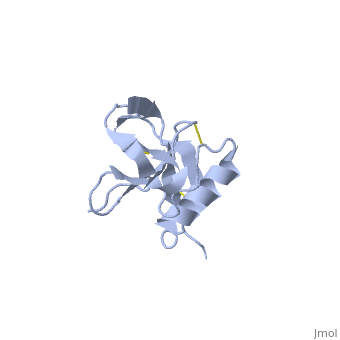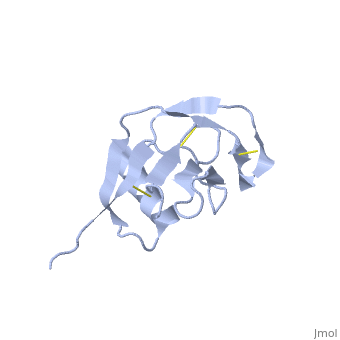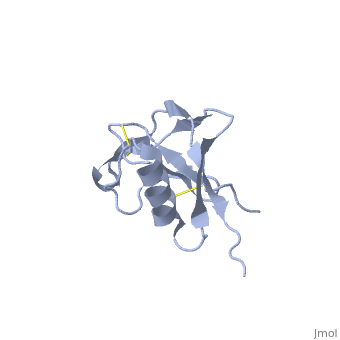
| - | + | <StructureSection load='1mwp' size='350' side='right' scene='' caption='Human amyloid precursor protein heparin-binding domain [[1mwp]]'> | |
== Function == | == Function == | ||
| Line 15: | Line 15: | ||
The extracellular region of APP contains several domains named E1 (residues 1-189) and E2 (residues 346-551). The E1 domain contains a growth factor-like or heparin-binding (residues 28-123) and Cu-binding (residues 124-189) subdomains. Additional domains are: Kunitz-type protease inhibitor (residues 287-344) and Zn-binding (residues 672-687). The Z-binding domain is involved in the oligomerizatin of APP. | The extracellular region of APP contains several domains named E1 (residues 1-189) and E2 (residues 346-551). The E1 domain contains a growth factor-like or heparin-binding (residues 28-123) and Cu-binding (residues 124-189) subdomains. Additional domains are: Kunitz-type protease inhibitor (residues 287-344) and Zn-binding (residues 672-687). The Z-binding domain is involved in the oligomerizatin of APP. | ||
| - | + | </StructureSection> | |
==3D structures of amyloid precursor protein== | ==3D structures of amyloid precursor protein== | ||
Revision as of 09:21, 7 December 2015
| |||||||||||
3D structures of amyloid precursor protein
Updated on 07-December-2015