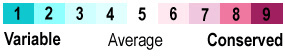This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Introduction to Evolutionary Conservation
From Proteopedia
(Difference between revisions)
| Line 99: | Line 99: | ||
===Sophisticated Analysis of Conservation: ConSurf=== | ===Sophisticated Analysis of Conservation: ConSurf=== | ||
| - | The above analysis assigns each amino acid in enolase to one of three categories: conserved, similar, or different. This is very simplistic. In contrast, the analysis used in Proteopedia (and in the above image) is sophisticated, using many more sequences, and weighting the impact of each sequence in the multiple sequence alignment according to the phylogenetic tree calculated from the alignment. This sophisticated determination of conservation and variability is done by the [[ConSurfDB vs. ConSurf|ConSurf Servers]] (see also summaries of their mechanism: [http://www.umass.edu/molvis/workshop/slides/consurf3.htm short version], or [[ConSurfDB_vs._ConSurf#The_ConSurf-DB_Mechanism|longer version]]). ConSurf divides conservation into 9 levels, and colors them as follows: | + | The above analysis assigns each amino acid in enolase to one of three categories: conserved, similar, or different. This is very simplistic, and sensitive to the addition or removal of one or a few sequences from the alignment which can have a large effect on the results. In contrast, the analysis used in Proteopedia (and in the above image) is sophisticated, using many more sequences, and weighting the impact of each sequence in the multiple sequence alignment according to the phylogenetic tree calculated from the alignment. This sophisticated determination of conservation and variability is done by the [[ConSurfDB vs. ConSurf|ConSurf Servers]] (see also summaries of their mechanism: [http://www.umass.edu/molvis/workshop/slides/consurf3.htm short version], or [[ConSurfDB_vs._ConSurf#The_ConSurf-DB_Mechanism|longer version]]). ConSurf's analysis is ''robust'': addition or removal of a few sequences has little effect. ConSurf divides conservation into 9 levels, and colors them as follows: |
<center>{{Template:ColorKey_ConSurf_NoGray}}</center> | <center>{{Template:ColorKey_ConSurf_NoGray}}</center> | ||
Revision as of 20:15, 27 February 2016
| |||||||||||
See Also
Notes and References
- ↑ MECP2 article in the National Library of Medicine's Genetic Home Reference
- ↑ Advantageous variability will be seen in these cases: 5hmg, 2vaa, 3hi6.