Sandbox Reserved 1172
From Proteopedia
(Difference between revisions)
| Line 11: | Line 11: | ||
=== Key Ligand Interactions === | === Key Ligand Interactions === | ||
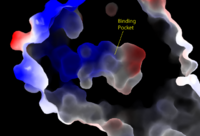
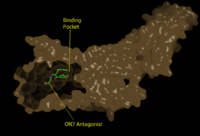
[[Image:Pic2proteo3.png|200 px|right|thumb|Figure 2: Surface representation of the LPA<sub>1</sub> receptor in tan interacting with its antagonist, ON7, shown in green and red sticks.The exterior of the protein was partially cut away to display the interior binding pocket.]] | [[Image:Pic2proteo3.png|200 px|right|thumb|Figure 2: Surface representation of the LPA<sub>1</sub> receptor in tan interacting with its antagonist, ON7, shown in green and red sticks.The exterior of the protein was partially cut away to display the interior binding pocket.]] | ||
| - | The biological ligand of the LPA<sub>1</sub> receptor receptor is lysophosphatidic acid, a phospholipid that contains a long, | + | The biological ligand of the LPA<sub>1</sub> receptor receptor is [https://en.wikipedia.org/wiki/Lysophosphatidic_acid lysophosphatidic acid (LPA)], a phospholipid that contains a long, nonpolar tail, a phosphate head, a chiral hydroxyl group, and an ester group. [http://www.guidetopharmacology.org/GRAC/LigandDisplayForward?ligandId=8589 ONO-9780307 (ON7)], is an antagonist for LPA due to its large nonpolar region, chiral hydroxyl group, ester, and carboxylic acid, which all resemble portions of the LPA molecule. The receptor provides specificity for its ligand by the amphipathic binding pocket; Three separate interactions with an antagonist of LPA<sub>1</sub>, help demonstrate the key interactions that stabilize the binding of the LPA phospholipid to this receptor. At the polar region of the binding pocket, the ligand binding is stabilized by <scene name='72/721543/Arg124gln125/3'>Arg124 and Gln125</scene> forming ionic and polar interactions with the carboxylic acid and the hydroxyl group of ON7.<ref name="regpeps">PMID: 26091040</ref> In addition, interplay between <scene name='72/721543/Lys39_and_glu293/7'>Glu293 and Lys39</scene> causes another stabilizing component with the ON7 antagonist. Glu293 forms polar interactions with Lys39, positioning it in close proximity to to the carboxylic acid of ON7, which then interactions with Lys39 via ionic bonding.<ref name="regpeps">PMID: 26091040</ref> While Lys39 is highly conserved among all six LPA receptors, a neighboring histidine residue is specific to the LPA<sub>1</sub> receptor. <scene name='72/721543/His40/1'>His40</scene> forms both ionic and polar interactions with the carboxylic acid of ON7. The protonation of this residue to greatly affects the binding affinity of LPA, and is an important link to tumor growth and survival in acidic environments.<ref name="regpeps">PMID: 26091040</ref> |
Revision as of 18:50, 14 April 2016
| This Sandbox is Reserved from Jan 11 through August 12, 2016 for use in the course CH462 Central Metabolism taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1160 through Sandbox Reserved 1184. |
To get started:
More help: Help:Editing |
Lysophosphatidic Acid Receptor 1
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 Chrencik JE, Roth CB, Terakado M, Kurata H, Omi R, Kihara Y, Warshaviak D, Nakade S, Asmar-Rovira G, Mileni M, Mizuno H, Griffith MT, Rodgers C, Han GW, Velasquez J, Chun J, Stevens RC, Hanson MA. Crystal Structure of Antagonist Bound Human Lysophosphatidic Acid Receptor 1. Cell. 2015 Jun 18;161(7):1633-43. doi: 10.1016/j.cell.2015.06.002. PMID:26091040 doi:http://dx.doi.org/10.1016/j.cell.2015.06.002
- ↑ Hernández-Méndez, Aurelio, Rocío Alcántara-Hernández, and J. Adolfo García-Sáinz. "Lysophosphatidic Acid LPA1-3 Receptors: Signaling, Regulation and in Silico Analysis of Their Putative Phosphorylation Sites." Receptors & Clinical Investigation Receptor Clin Invest 1.3 (2014). Web. 15 Feb. 2016.'
- ↑ Yung, Y. C., N. C. Stoddard, and J. Chun. "LPA Receptor Signaling: Pharmacology, Physiology, and Pathophysiology." The Journal of Lipid Research 55.7 (2014): 1192-214. Web. 17 Feb. 2016.'
- ↑ Chun, J., Hla, T., Spiegel, S., and Moolenaar, W.H. “Lysophospholipid Receptors: Signaling and Biochemistry.” John Wiley & Sons, Inc. (2013) pp.i-xviii. 5 Feb. 2016.'
- ↑ Anliker B, Choi JW, Lin ME, Gardell SE, Rivera RR, Kennedy G, Chun J. Lysophosphatidic acid (LPA) and its receptor, LPA1 , influence embryonic schwann cell migration, myelination, and cell-to-axon segregation. Glia. 2013 Dec;61(12):2009-22. doi: 10.1002/glia.22572. Epub 2013 Sep 24. PMID:24115248 doi:http://dx.doi.org/10.1002/glia.22572
- ↑ Chun, E., Thompson, A.A., Lui, W., Roth, C.B., Griffith, M.T., Katritch, V., Kunken, J., Xu, F., Cherezov, V., Hanson, M.A., and Stevens, R.C. “Fusion partner tool chest for the stabilization and crystallization of G protein-coupled receptors.” Structure 20, (2012) 967-976.'
- ↑ Van Durme, J., Horn, F., Costagliola, S., Vriend, G., and Vassart, G. “GRIS: glycoprotein-hormone receptor information system.” Mol. (2006) Endocrinol. 20, 2247-2255'
- ↑ Lin ME, Herr DR, Chun J. Lysophosphatidic acid (LPA) receptors: signaling properties and disease relevance. Prostaglandins Other Lipid Mediat. 2010 Apr;91(3-4):130-8. doi:, 10.1016/j.prostaglandins.2009.02.002. Epub 2009 Mar 4. PMID:20331961 doi:http://dx.doi.org/10.1016/j.prostaglandins.2009.02.002
- ↑ Justus CR, Dong L, Yang LV. Acidic tumor microenvironment and pH-sensing G protein-coupled receptors. Front Physiol. 2013 Dec 5;4:354. doi: 10.3389/fphys.2013.00354. PMID:24367336 doi:http://dx.doi.org/10.3389/fphys.2013.00354