This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Rhomboid protease
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Structural highlights == | == Structural highlights == | ||
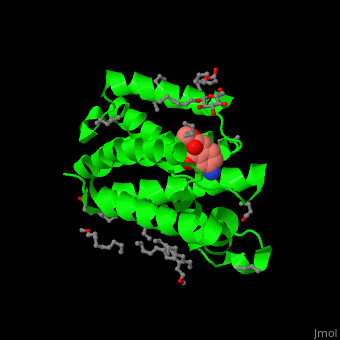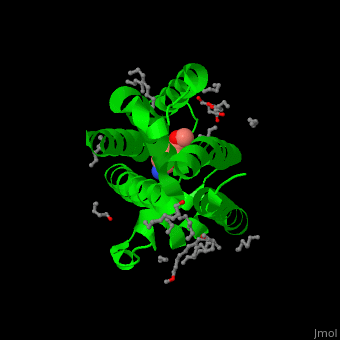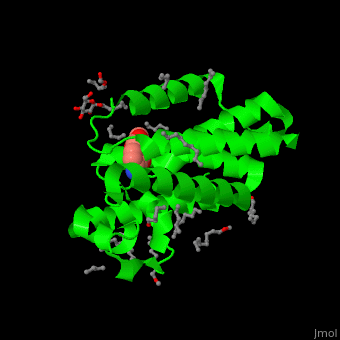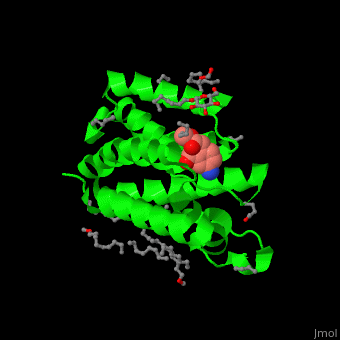
| - | The inhibitor isocoumarin binds to the GlpG catalytic residues Ser and His<ref>PMID:20890268</ref>. | + | The inhibitor <scene name='55/551211/Cv/3'>isocoumarin covalently binds to the GlpG catalytic residues Ser and His</scene> (in deepskyblue)<ref>PMID:20890268</ref>. <scene name='55/551211/Cv/4'>Whole active site</scene>. Water molecules shown as red spheres. |
</StructureSection> | </StructureSection> | ||
Revision as of 11:18, 10 August 2016
| |||||||||||
3D Structures of rhomboid protease
Updated on 10-August-2016
References
- ↑ Freeman M. Rhomboid proteases and their biological functions. Annu Rev Genet. 2008;42:191-210. doi: 10.1146/annurev.genet.42.110807.091628. PMID:18605900 doi:http://dx.doi.org/10.1146/annurev.genet.42.110807.091628
- ↑ Vinothkumar KR, Strisovsky K, Andreeva A, Christova Y, Verhelst S, Freeman M. The structural basis for catalysis and substrate specificity of a rhomboid protease. EMBO J. 2010 Oct 1. PMID:20890268 doi:10.1038/emboj.2010.243