Immunodeficiency virus protease
From Proteopedia
| Line 18: | Line 18: | ||
==Structure of HIV-1 Protease== | ==Structure of HIV-1 Protease== | ||
The X-ray structure of HIV-1 protease<ref>PMID:2548279</ref><ref>PMID:2682266</ref> reveals that it is composed of <scene name='User:David_Canner/Sandbox_HIV/Identical_subunits/1'>two symmetrically related subunits</scene>, each consisting of 99 amino acid residues. The subunits come together in such as way as to <scene name='User:David_Canner/Sandbox_HIV/Tunnel/1'>form a tunnel where they meet</scene>. This tunnel is of critical importance because the active site of the protease is located in its interior. The active site consists of <scene name='User:David_Canner/Sandbox_HIV/Catalytic_triad/3'> two Asp-Thr-Gly conserved sequences</scene>, making it a member of the aspartyl protease family. The two Asp's are <scene name='User:David_Canner/Sandbox_HIV/Catalytic_asp/1'>essential catalytic residues</scene> either interact with the incoming water OR protonate the carbonyl to make the carbon more electrophilic for the incoming <scene name='31/315240/Saquinavir_cat_water/2'>water</scene>. You may be wondering how a polyprotein makes its way into the active-site tunnel, as the<scene name='User:David_Canner/Sandbox_HIV/Narrow_tunnel/1'> tunnel appears to be too narrow </scene> to admit it. The key is the two flexible flaps on the top of the tunnel that <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph/3'>move to allow proteins </scene>to enter the tunnel. The flaps <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph_flaps/2'>undergo a dramatic movement</scene>, shifting from an open to a closed conformation to bind the target in an appropriate conformation for cleavage. | The X-ray structure of HIV-1 protease<ref>PMID:2548279</ref><ref>PMID:2682266</ref> reveals that it is composed of <scene name='User:David_Canner/Sandbox_HIV/Identical_subunits/1'>two symmetrically related subunits</scene>, each consisting of 99 amino acid residues. The subunits come together in such as way as to <scene name='User:David_Canner/Sandbox_HIV/Tunnel/1'>form a tunnel where they meet</scene>. This tunnel is of critical importance because the active site of the protease is located in its interior. The active site consists of <scene name='User:David_Canner/Sandbox_HIV/Catalytic_triad/3'> two Asp-Thr-Gly conserved sequences</scene>, making it a member of the aspartyl protease family. The two Asp's are <scene name='User:David_Canner/Sandbox_HIV/Catalytic_asp/1'>essential catalytic residues</scene> either interact with the incoming water OR protonate the carbonyl to make the carbon more electrophilic for the incoming <scene name='31/315240/Saquinavir_cat_water/2'>water</scene>. You may be wondering how a polyprotein makes its way into the active-site tunnel, as the<scene name='User:David_Canner/Sandbox_HIV/Narrow_tunnel/1'> tunnel appears to be too narrow </scene> to admit it. The key is the two flexible flaps on the top of the tunnel that <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph/3'>move to allow proteins </scene>to enter the tunnel. The flaps <scene name='User:David_Canner/Sandbox_HIV/Hiv_tunnel_morph_flaps/2'>undergo a dramatic movement</scene>, shifting from an open to a closed conformation to bind the target in an appropriate conformation for cleavage. | ||
| - | <br> | ||
| - | <br> HIV Protease Movie by Warren L. DeLano (made via PyMol) | ||
| - | <html5media height=“500” width=“1000”>https://www.youtube.com/embed/iSeVYYDvLCk</html5media> | ||
==Medical Implications== | ==Medical Implications== | ||
There currently is no cure or vaccine against HIV. Researchers, however, have discovered treatments that can halt and even reverse the progression of AIDS, due in large part to our understanding of the structure of HIV-1 protease. <scene name='User:David_Canner/Sandbox_HIV/Saquinavir/4'>Saquinavir</scene> ([[Invirase]]) was the first protease inhibitor approved by the FDA for the treatment of HIV. It inhibits HIV protease by <scene name='User:David_Canner/Sandbox_HIV/Saquinavir_tunnel/1'>binding tightly in the active site tunnel</scene>, preventing the binding of polyproteins. Its chemical structure mimics the tetrahedral intermediate of the hydrolytic reaction, thereby <scene name='User:David_Canner/Sandbox_HIV/Saquinavir_cat/3'>interacting strongly with the catalytic Asp residues</scene>.<ref>PMID:17243183</ref> Saquinavir is essentially an uncleavable ligand, as indicated by the <scene name='User:David_Canner/Sandbox_HIV/Hiv_morph2/9'> similar conformational changes in the protease flaps </scene> on binding saquinavir or a polypeptide . Resistance to saquinavir is due to alterations in the HIV protease sequence, including the mutation of <scene name='31/315240/Saquinavir_mut/1'>Leu 10 and Ile 50</scene><ref>PMID: 8969180</ref>. Drugs used to treat HIV infection that inhibit <scene name='User:David_Canner/Sandbox_HIV/Inhibitor_intro/1'>HIV protease</scene> include <scene name='User:David_Canner/Sandbox_HIV/Indinavir/2'>Indinavir </scene> ([[Crixivan]]), <scene name='User:David_Canner/Sandbox_HIV/Ritonavir/1'>Ritonavir</scene> ([[Norvir]]), [[Saquinavir]], [[Tipranavir]], [[Amprenavir]] (Agenerase), [[Atazanavir]] (Rayataz), [[Darunavir]] (Prezista), [[Fosamprenavir]] (Lexiva or Telzir), [[Lopinavir]] (Kaletra), [[Nelfinavir]] (Viracept) and <scene name='User:David_Canner/Sandbox_HIV/Nelfinavir/2'>Nelfinavir</scene> ([[Viracept]]). | There currently is no cure or vaccine against HIV. Researchers, however, have discovered treatments that can halt and even reverse the progression of AIDS, due in large part to our understanding of the structure of HIV-1 protease. <scene name='User:David_Canner/Sandbox_HIV/Saquinavir/4'>Saquinavir</scene> ([[Invirase]]) was the first protease inhibitor approved by the FDA for the treatment of HIV. It inhibits HIV protease by <scene name='User:David_Canner/Sandbox_HIV/Saquinavir_tunnel/1'>binding tightly in the active site tunnel</scene>, preventing the binding of polyproteins. Its chemical structure mimics the tetrahedral intermediate of the hydrolytic reaction, thereby <scene name='User:David_Canner/Sandbox_HIV/Saquinavir_cat/3'>interacting strongly with the catalytic Asp residues</scene>.<ref>PMID:17243183</ref> Saquinavir is essentially an uncleavable ligand, as indicated by the <scene name='User:David_Canner/Sandbox_HIV/Hiv_morph2/9'> similar conformational changes in the protease flaps </scene> on binding saquinavir or a polypeptide . Resistance to saquinavir is due to alterations in the HIV protease sequence, including the mutation of <scene name='31/315240/Saquinavir_mut/1'>Leu 10 and Ile 50</scene><ref>PMID: 8969180</ref>. Drugs used to treat HIV infection that inhibit <scene name='User:David_Canner/Sandbox_HIV/Inhibitor_intro/1'>HIV protease</scene> include <scene name='User:David_Canner/Sandbox_HIV/Indinavir/2'>Indinavir </scene> ([[Crixivan]]), <scene name='User:David_Canner/Sandbox_HIV/Ritonavir/1'>Ritonavir</scene> ([[Norvir]]), [[Saquinavir]], [[Tipranavir]], [[Amprenavir]] (Agenerase), [[Atazanavir]] (Rayataz), [[Darunavir]] (Prezista), [[Fosamprenavir]] (Lexiva or Telzir), [[Lopinavir]] (Kaletra), [[Nelfinavir]] (Viracept) and <scene name='User:David_Canner/Sandbox_HIV/Nelfinavir/2'>Nelfinavir</scene> ([[Viracept]]). | ||
| + | |||
| + | ==HIV Protease Movie== | ||
| + | <br> | ||
| + | <br> HIV Protease Movie by Warren L. DeLano (made via PyMol) | ||
| + | <html5media height=“500” width=“1000”>https://www.youtube.com/embed/iSeVYYDvLCk</html5media> | ||
== Structural Insights into the South African HIV-1 Subtype C Protease: Impact of hinge region dynamics and flap flexibility in drug resistance <ref>doi 10.1080/07391102.2012.736774</ref>== | == Structural Insights into the South African HIV-1 Subtype C Protease: Impact of hinge region dynamics and flap flexibility in drug resistance <ref>doi 10.1080/07391102.2012.736774</ref>== | ||
Revision as of 11:42, 29 May 2018
| |||||||||||
Contents |
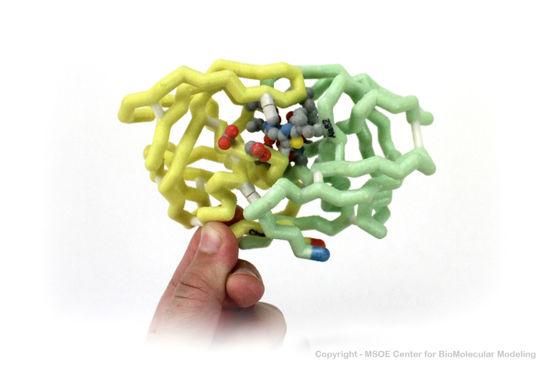
3D Printed Physical Model of HIV Protease
Shown below are 3D printed physical models of HIV Protease. Both versions are shown in alpha carbon format, with select side chains shown colored by element, with carbon gray, nitrogen blue, oxygen red and sulfur yellow. Both models have been designed with precisely embedded magnets that allow the two chains to pull apart into individual pieces.
The MSOE Center for BioMolecular Modeling
The MSOE Center for BioMolecular Modeling uses 3D printing technology to create physical models of protein and molecular structures, making the invisible molecular world more tangible and comprehensible. To view more protein structure models, visit our Model Gallery.
3D structures of immunodeficiency protease
Updated on 29-May-2018
Additional Resources
For additional information, see:
- Human Immunodeficiency Virus
- Structural Biology of HIV, an interactive Flash graphic of the virion with explanations of its components.
References
- ↑ Tie Y, Wang YF, Boross PI, Chiu TY, Ghosh AK, Tozser J, Louis JM, Harrison RW, Weber IT. Critical differences in HIV-1 and HIV-2 protease specificity for clinical inhibitors. Protein Sci. 2012 Mar;21(3):339-50. doi: 10.1002/pro.2019. Epub 2012 Jan 24. PMID:22238126 doi:10.1002/pro.2019
- ↑ Wlodawer A, Miller M, Jaskolski M, Sathyanarayana BK, Baldwin E, Weber IT, Selk LM, Clawson L, Schneider J, Kent SB. Conserved folding in retroviral proteases: crystal structure of a synthetic HIV-1 protease. Science. 1989 Aug 11;245(4918):616-21. PMID:2548279
- ↑ Lapatto R, Blundell T, Hemmings A, Overington J, Wilderspin A, Wood S, Merson JR, Whittle PJ, Danley DE, Geoghegan KF, et al.. X-ray analysis of HIV-1 proteinase at 2.7 A resolution confirms structural homology among retroviral enzymes. Nature. 1989 Nov 16;342(6247):299-302. PMID:2682266 doi:http://dx.doi.org/10.1038/342299a0
- ↑ Tie Y, Kovalevsky AY, Boross P, Wang YF, Ghosh AK, Tozser J, Harrison RW, Weber IT. Atomic resolution crystal structures of HIV-1 protease and mutants V82A and I84V with saquinavir. Proteins. 2007 Apr 1;67(1):232-42. PMID:17243183 doi:10.1002/prot.21304
- ↑ Maschera B, Darby G, Palu G, Wright LL, Tisdale M, Myers R, Blair ED, Furfine ES. Human immunodeficiency virus. Mutations in the viral protease that confer resistance to saquinavir increase the dissociation rate constant of the protease-saquinavir complex. J Biol Chem. 1996 Dec 27;271(52):33231-5. PMID:8969180
- ↑ Naicker P, Achilonu I, Fanucchi S, Fernandes M, Ibrahim MA, Dirr HW, Soliman ME, Sayed Y. Structural insights into the South African HIV-1 subtype C protease: impact of hinge region dynamics and flap flexibility in drug resistance. J Biomol Struct Dyn. 2012 Nov 12. PMID:23140382 doi:10.1080/07391102.2012.736774
Proteopedia Page Contributors and Editors (what is this?)
Joel L. Sussman, Michal Harel, Eran Hodis, Mark Hoelzer, David Canner, Eric Martz, Ann Taylor, Wayne Decatur, Alexander Berchansky, Jaime Prilusky, Karsten Theis