This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1506
From Proteopedia
| Line 1: | Line 1: | ||
| - | <Structure load=' | + | <Structure load='Insert PDB code or filename here' size='350' frame='true' align='right' caption='Insert caption here' scene='Insert optional scene name here' /><scene name='80/802680/Anaat/1'>ANAAT en couleur</scene> |
'<Structure load='1b6b' size='350' frame='true' align='right' caption='Protein Display ' scene='Insert optional scene name here' />'{{Sandbox_Reserved_ESBS}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> | '<Structure load='1b6b' size='350' frame='true' align='right' caption='Protein Display ' scene='Insert optional scene name here' />'{{Sandbox_Reserved_ESBS}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> | ||
== Structure == | == Structure == | ||
| Line 7: | Line 7: | ||
== Function == | == Function == | ||
| + | Serotonin N-acetyltransferase is also named aralkylamine-N-acetyltransferase (ANAAT). | ||
| + | [[Image:melationin synthesis.jpg]] | ||
== Regulation == | == Regulation == | ||
| Line 16: | Line 18: | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
| + | [1] The Structural Basis of Ordered Substrate Binding by Serotonin N-Acetyltransferase: Enzyme Complex at 1.8 A ˚ Resolution with a Bisubstrate Analog - Alison Burgess Hickman,M. A. A. Namboodiri, David C. Klein, and Fred Dyda - Cell, Vol. 97, 361–369, April 30, 1999. | ||
| + | |||
| + | [2] Melatonin synthesis: 14-3-3-dependent activation and inhibition of arylalkylamine N -acetyltransferase mediated by phosphoserine-205 - Surajit Ganguly, Joan L. Weller, Anthony Ho, Philippe Chemineau, Benoit Malpaux, and David C. Klein - PNAS, January 25, 2005. | ||
| + | |||
| + | [3] | ||
| + | |||
<references/> | <references/> | ||
Revision as of 11:33, 2 January 2019
|
|
| This Sandbox is Reserved from 06/12/2018, through 30/06/2019 for use in the course "Structural Biology" taught by Bruno Kieffer at the University of Strasbourg, ESBS. This reservation includes Sandbox Reserved 1480 through Sandbox Reserved 1543. |
To get started:
More help: Help:Editing |
Contents |
Structure
This is a default text for your page b6. Click above on edit this page to modify. Be careful with the < and > signs. You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Function
Serotonin N-acetyltransferase is also named aralkylamine-N-acetyltransferase (ANAAT).
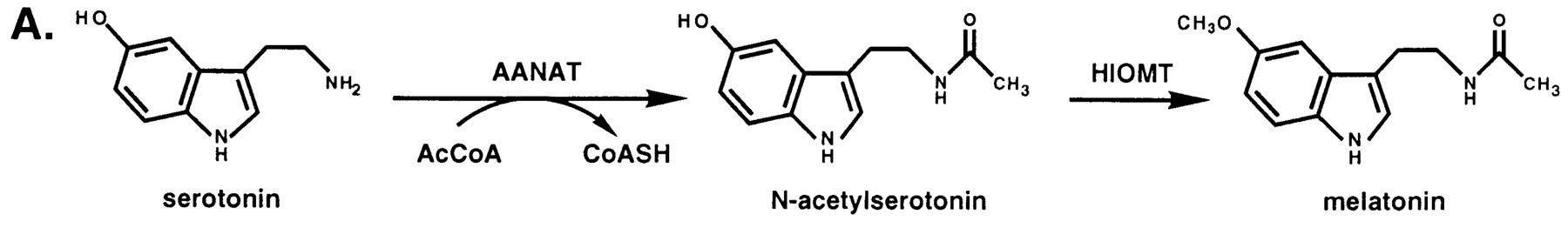
Regulation
Relevance
Structural highlights
</StructureSection>
References
[1] The Structural Basis of Ordered Substrate Binding by Serotonin N-Acetyltransferase: Enzyme Complex at 1.8 A ˚ Resolution with a Bisubstrate Analog - Alison Burgess Hickman,M. A. A. Namboodiri, David C. Klein, and Fred Dyda - Cell, Vol. 97, 361–369, April 30, 1999.
[2] Melatonin synthesis: 14-3-3-dependent activation and inhibition of arylalkylamine N -acetyltransferase mediated by phosphoserine-205 - Surajit Ganguly, Joan L. Weller, Anthony Ho, Philippe Chemineau, Benoit Malpaux, and David C. Klein - PNAS, January 25, 2005.
[3]
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
