This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Ashley Crotteau/Sandbox1
From Proteopedia
(Difference between revisions)
| Line 39: | Line 39: | ||
===Renal Fibrosis=== | ===Renal Fibrosis=== | ||
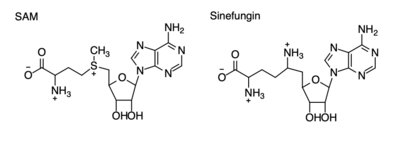
| - | [[Image: | + | [[Image:Sam_vs_sinefungin.png|400 px|right|thumb|Figure 4: Structural differences between SAM and Sinefungin]] |
The inhibition of KMT SET 7/9 has been found to enhance renal fibrosis.<ref name="Sun" /> H3K4 methylation activates the transcription of fibrotic genes, and the suppression of the H3K4 methylation was found to enhance renal fibrosis in a mouse model.<ref name="Sun" /> Sinefungin, a competitive methyltransferase inhibitor, binds to KMT to inhibit SAM.<ref name="Sun" /> SAM and Sinefungin have similar structures (Figure 4), differing only with the removal of a sulfide and the replacement of a methyl to an amine. The amine replaces the methyl donor for the reaction of KMT, inhibiting the reaction and preventing methylation. Without the methylation of H3K4, the transcription of fibrotic genes is deactivated, leading to renal fibrosis.<ref name="Sun" /> | The inhibition of KMT SET 7/9 has been found to enhance renal fibrosis.<ref name="Sun" /> H3K4 methylation activates the transcription of fibrotic genes, and the suppression of the H3K4 methylation was found to enhance renal fibrosis in a mouse model.<ref name="Sun" /> Sinefungin, a competitive methyltransferase inhibitor, binds to KMT to inhibit SAM.<ref name="Sun" /> SAM and Sinefungin have similar structures (Figure 4), differing only with the removal of a sulfide and the replacement of a methyl to an amine. The amine replaces the methyl donor for the reaction of KMT, inhibiting the reaction and preventing methylation. Without the methylation of H3K4, the transcription of fibrotic genes is deactivated, leading to renal fibrosis.<ref name="Sun" /> | ||
Revision as of 18:38, 5 April 2019
H. sapiens Lysine Methyltransferase, SET 7/9
| |||||||||||
References
[3] [5] [1] [2] [4] [8] [6] [7]
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 Schubert HL, Blumenthal RM, Cheng X. Many paths to methyltransfer: a chronicle of convergence. Trends Biochem Sci. 2003 Jun;28(6):329-35. PMID:12826405
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Yeates TO. Structures of SET domain proteins: protein lysine methyltransferases make their mark. Cell. 2002 Oct 4;111(1):5-7. PMID:12372294
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378
- ↑ 4.0 4.1 Huang S, Shao G, Liu L. The PR domain of the Rb-binding zinc finger protein RIZ1 is a protein binding interface and is related to the SET domain functioning in chromatin-mediated gene expression. J Biol Chem. 1998 Jun 26;273(26):15933-9. PMID:9632640
- ↑ 5.0 5.1 doi: https://dx.doi.org/10.1016/C2014-0-02189-2
- ↑ 6.0 6.1 6.2 6.3 6.4 6.5 Del Rizzo PA, Couture JF, Dirk LM, Strunk BS, Roiko MS, Brunzelle JS, Houtz RL, Trievel RC. SET7/9 catalytic mutants reveal the role of active site water molecules in lysine multiple methylation. J Biol Chem. 2010 Oct 8;285(41):31849-58. Epub 2010 Aug 1. PMID:20675860 doi:http://dx.doi.org/10.1074/jbc.M110.114587
- ↑ 7.0 7.1 7.2 7.3 7.4 Sun G, Reddy MA, Yuan H, Lanting L, Kato M, Natarajan R. Epigenetic histone methylation modulates fibrotic gene expression. J Am Soc Nephrol. 2010 Dec;21(12):2069-80. doi: 10.1681/ASN.2010060633. Epub 2010, Oct 7. PMID:20930066 doi:http://dx.doi.org/10.1681/ASN.2010060633
- ↑ https://en.wikipedia.org/wiki/SET_domain#Structure
Student Contributors
Ashley Crotteau
Parker Hiday
Lauren Allman